Modeling protein motors
340 likes | 453 Views
Explore theoretical models and simulation frameworks for protein motors, including mesoscopic models and kinetic models. Dive into the bacterial flagellar motor and F1Fo ATPase, understanding torque-speed relationships and mechanics. Learn how chemical energy powers molecular motors and the construction of mesoscopic models. Discover the physical explanations behind torque-speed relations and how to fit models to experimental data.

Modeling protein motors
E N D
Presentation Transcript
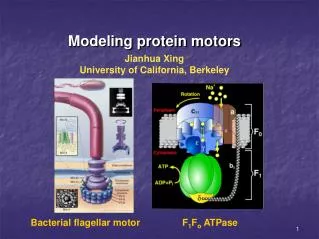
Modeling protein motors Jianhua Xing University of California, Berkeley Bacterial flagellar motor F1Fo ATPase
Question 1: What are appropriate theoretical and simulation frameworks? A theoretical model should be • Simple • Not over-simplified • Useful
Simple • Limited information • Insufficient mechanics treatment Mesoscopic models Chemical states of a rotor binding site: E: empty O: occupied • Computationally less demanding than atomic level simulations • Treats mechanics appropriately • Atomic level simulations • Provide atomic details • Limited by system size and simulation length • Require detailed structures Kinetic models
+ x _ I. General discussion 1. Molecular motors use chemical energy to perform mechanical work Occupied Potential Empty + Occupied state Empty state x Chemical transitions switch the system from one potential curve to another, and result in mechanical motion
Dq DG rotor q Dq stator A generic molecular motor Free energy Angular position q Keller, Bustamante (2000), Biophys. J., 78: 541-556
II. The Bacteria Flagellar motor 1. Summary of experimental studies H. Berg’s website
Vibrio alginolyticus a). Structure of bacterial flagellar motor D. Thomas, N. Francis, and D. Derosier, unpublished EM image
periplasm cytoplasm pmf = proton motive force ion = proton, sodium The “fuels”: trans-membrane ion motive force
b) Mechano-chemical studies • The flagellar motor can • Generate torque 2700 ~ 4000 pN.nm • Rotate as fast as 1700 Hz • (1 Hz = 1 cycle/second) Under viscous load Under rotating electric field Fung, Berg (1995), Nature, 375: 809-812 Ryu, Berry, Berg (2000), Nature, 403:444-447 Chen, Berg (2000), Biophys. J. 78: 1036-1041 Gabel, Berg (2003), PNAS, 100:8748-8751 Sowa et al. (2005), Nature, 437: 916-919
c) The Puzzle: ~ Linear decrease Torque ~ Constant Speed pmf Fung, Berg (1995), Nature, 375: 809-812 Ryu, Berry, Berg (2000), Nature, 403:444-447 Chen, Berg (2000), Biophys. J. 78: 1036-1041 Gabel, Berg (2003), PNAS, 100:8748-8751
2. Our model a) We proposed the working mechanism based on available information Xing, Bai, Berry , Oster (2005), submitted
b) We identified key ingredients from experiments The power stroke is driven by a conformational transformation in the stator that is triggered by the protons hopping onto and off the stator Elastic coupling Tight coupling The rotation of the motor is observed through a soft elastic linkage between the motor and the viscous load The ion channel through the stator is gated by the motion of the rotor Xing, Bai, Berry , Oster (2005), submitted
Prob. density c) Mathematical modeling Motor Torque Brownian Torque Markovian chemical transitions Viscous Torque Load Torque + Langevin Fokker-Planck Potential based kinetic model Wang, Peskin, Elston (2003), J. Theor. Biol. 221:491-511 Xing, Wang, Oster (2005), Biophys. J. 89:1551-1563
d) Our model can fit the data well Xing, Bai, Berry, Oster (2005), submitted
3. Physical explanation of the torque-speed relation a) Plateau: time scale separation & soft linkage Spring constant
bead DG/Dq motor Crossover between the two time scales
Elastic linkage allows multiple time-scales Conservative load Viscous load with spring Viscous load with rigid linkage Xing, Bai, Berry, Oster (2005), submitted
Pushed away from transition region Free energy Angular position q Free energy Angular position q b) Sharp transition: positive feedback
rotor Ryu, Berry, Berg (2000), Nature, 403:444-447 Low speed region: thermodynamic limit c) Prediction more stators more power High speed region: Destructive interference Xing, Bai, Berry, Oster (2005), submitted
more stators more steps: depends on relative phases between stators Free energy Angular position q 2 stators 1 stator
d) The same physics explains other puzzles Yasuda et al. (1998), Cell (93) 1117-1124 Mycoplasma causes diseases like pneumonia. Its sliding rate depends on temperature linearly. Miyata, Ryu, Berg (2002), J. Bacter., 1827-1131
4. Summary of flagellar motor • The observed flagellar motor dynamics is due to • Internal conformational change • Soft elastic linkage between motor and load • Tight coupling • Localized chemical transitions
Dynamics Structure Mathematical modeling Question 2: How to construct a mesoscopic model? Fit data and suggest experiments
III. ATP synthesis with the F1Fo ATPase 1. the F1Fo ATPase THE F1Fo ATPase THE F1 PART Fo model: F1 model: Xing, et al. (2004), Biophys. J. 87: 2148-2163 Xing, Liao, Oster (2005), PNAS, in press
F1 Fo 2. The F1 part Various experimental studies serve as the model basis Single molecule measurements Biochemistry measurements Crystal structures Mutation studies
Intrinsic elastic energy open close open Empty ADP + Pi ATP Free energy open closed Empty ~ 20 KBT ADP 200 300 0 100 Substrate binding energy b/g & b/e interactions Rotation angle [degree] Constructing the potentials: qualitative thinking Berzborn, Schlittler (2002), FEBS Lett. 26828, 1-8 Ma, et al. (2002), Structure 10, 921-931
Fo torque: Empty open closed Constructing the potentials: qualitative thinking open close open Rotation without synthesis Correct sequence Free energy G ADP+Pi ATP ADP Empty 200 300 0 100 Rotation angle [degree] Xing, Liao, Oster (2005), PNAS, in press
Empty ADP Right timing/tight coupling is ensured by specific b/g & b/e interactions Xing, Liao, Oster (2005), PNAS, in press
Boyer’s binding change mechanism From announcement of 1997 Nobel Prize in chemistry ~ 20 KBT Require pmf ADP binding helps ATP release at another site Mg.ADP inhibition state Reveal dynamic features withoutany calculation! Free energy G Xing, Liao, Oster (2005), PNAS, in press
The model can fit steady state ATP synthesis data The model reveals multiple reaction pathways Tomashek, JJ et al. (2004), J. Biol. Chem. 279: 4465-4470 Xing, Liao, Oster (2005), PNAS, in press
3. Summary of the model on ATP synthesis • Our model • integrates a large body of experimental observations • proposes a set of free energy potentials, which contains almost all dynamic information • makes many experimentally testable predictions
IV. SUMMARY Theoretical/Computational/Systems biology: require intimate collaboration between experimentalists & theoretician I have used molecular motor studies to show that theoretical modeling can: • Integrate and transform information • Bridge structural and dynamical studies • Predict dynamical behaviors • Suggest new experiments
V. ACKNOWLEDGMENT • Richard Berry (Oxford) • Fan Bai (Oxford) • Peter Dimroth (ETH, Switzerland) • Christoph v. Balmoos (ETH, Switzerland) • George Oster • Jung-Chi Liao • Oleg Igoshin • Andrew Spakowitz • Oleksii Sliusarenko • Jing Chen • Joshua Adelman • Hongyun Wang (UC Santa Cruz) • Sean Sun (JHU) Financial support: NIH
δ a c11 Fo b γ ε F1 α3β3 δ The End