Molecular Genetics - Biotechnology
310 likes | 504 Views
Molecular Genetics - Biotechnology. Chapter 6. Tools and Techniques – 6.1.

Molecular Genetics - Biotechnology
E N D
Presentation Transcript
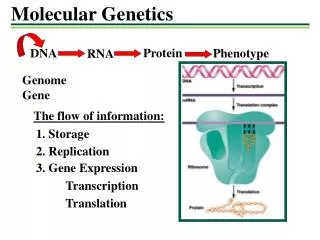
Molecular Genetics - Biotechnology Chapter 6
Tools and Techniques – 6.1 • The United Nations Convention on Biological Diversity defines biotechnology as: "Biotechnology" means any technological application that uses biological systems, living organisms, or derivatives thereof, to make or modify products or processes for specific use." • Specialized tools that need to be used by molecular biologists to allow them to work with DNA. • Recombinant DNA fragment of DNA composed of sequences originating from at least two different sources
Tools and Techniques • Restriction Endonucleases enzymes that are like molecular scissors and cut DNA at specific base pair sequences (restriction enzymes). The cut is a hydrolysis reaction at the phosphodiester bonds • The site where the restriction enzyme works is called a recognition site and it is usually palindromic and consists of four to eight base pairs • Sticky ends fragment end of a DNA molecule with short single stranded overhangs; EcoRI • More useful to molecular biologists as H bonds start to anneal naturally
Tools and Techniques • Blunt ends fragment end of a DNA molecule with bases fully paired (flush end); SmaI • Usefulness EcoRI has a six base pair recognition site (occurs once in every 4096 nucleotides = 46) • not too many cuts to destroy whole genes but enough cuts to make the DNA workable
Tools and Techniques • Naming of Restriction Enzymes • BamHI • B genus Bacillus • am species amyloliquefaciens • H strain • I first endonuclease isolated from the strain
Tools and Techniques • Methylases & DNA Ligase • Methylases enzymes that add a methyl group to one of the nucleotides found in a restriction endonuclease recognition site ensuring that the prokaryotes own DNA does not get cleaved • DNA Ligase an enzyme used to join together the phosphodiester backbone of DNA via a condensation reaction • T4 DNA Ligase enzyme from T4 bacteriophage that joins blunt ends together
Tools and Techniques • Gel electrophoresis separation of charged molecules on the basis of size taking advantage of the chemical and physical properties of DNA • DNA is negatively charged each nucleotide carries a phosphate group that has a charge of -1 • A charge is passed through the electrolyte buffer solution with the negative end where the DNA is loaded to the opposite positive end
Tools and Techniques • DNA can be cut with restriction enzymes migration through the gel is inversely proportional to the logarithm of their size…that is, the shorter pieces move further, the longer pieces do not move very far • DNA is mixed with a loading dye (visualization of DNA) and glycerol (weighs DNA down so it will fall into the wells).
Tools and Techniques • Gel is made from agarose (seaweed polysaccharide) or polyacrylamide (artificial polymer) • Gel gets stained with ethidium bromide that incorporates itself into the rungs of the DNA molecules and fluoresces under UV light.
Tools and Techniques • Bacteria often provide the appropriate machinery (enzymes and ribosomes) for us to produce proteins from a specific gene insulin example • Bacteria have small circular pieces of DNA called plasmids within their cytoplasm and the relationship is endosymbiotic. Number of base pairs ranges from 1000 – 200 000. • The number of plasmids in a bacteria is known as the copy number; the more plasmids, the more protein that can be created
Tools and Techniques • Transformation introduction of foreign DNA, by a plasmid or virus, into a bacterial cell • Plasmids can be vectors that carry DNA to be introduced into host cells
Tools and Techniques • Calcium Chloride Method • Bacterial cells are suspended in 0oC, making the cell membrane more rigid. • Bacterial membrane contains exposed phosphates which are naturally negatively charged. • Positively charged calcium ions stabilize the negative phosphates. • Plasmid DNA is now introduced as the membrane is chemically and physically “frozen”. • The negatively charged phosphates of the DNA are stabilized by the calcium cations • Shock treatment of 42oC for 90 seconds which causes the outside of the cell to be warmer than the inside. • This creates a draft which sweeps the plasmid into the bacterial cell through its porous membrane. • Environment goes back to 37oC.
Tools and Techniques • Electroporators – chambers that subject bacteria to an electric shock which loosens the structure of cell walls and allows DNA to enter host cells. • Gene guns – water or He to shoot towards DNA wrapped on a gold particle. The force of the impact causes the DNA to be propelled toward the target cells where it penetrates cell walls and membranes. • Biolistic Particle Delivery System
Tools and Techniques • Selective plating allows you to see if the process has worked. • Attempt to grow new plasmid in an antibiotic environment. If it grows, then you know the plasmid was introduced as the plasmid not only had the gene of desire in it, but also an antibiotic resistant gene in it.
Genetic Engineering – 6.2 • Genetic Engineering altering the sequence of DNA molecules • First completed in the early 1970s by Cohen (plasmids) and Boyer (restriction endonucleases) • Insulin 90% of diabetics get this protein from bacterial genetic engineering • Somatropin similar to growth hormone somatotropin which is used to treat dwarfism and Turner’s syndrome • Also used to treat AIDS associated wasting syndrome as it builds muscle
Advanced Techniques - 6.3 • Polymerase Chain Reaction – Mullis 1987 (p. 297, fig. 1) • PCR exponential amplification of DNA sequence by repeated cycles of strand separation and replication. • The methodology is closely related to DNA replication: • Two strands of DNA separated using DNA gyrase and DNA helicase.
Advanced Techniques - 6.3 • To further facilitate the separation heat is used (94oC-96oC) H bonds get broken easily. • Once separate the individual strands can be used as template strands. • DNA primers are used in place of RNA primers (DNA replication process) as DNA primers can be easily created in laboratory settings. • They are complementary to the opposing 3’-5’ends to the DNA target sequence to be replicated. • The temperature is dropped to 50oC-65oC for the primers to anneal with the template DNA.
Advanced Techniques • Taq polymerase (a high temperature DNA polymerase) adds complementary nucleotides. • Process takes place at 72oC which Taq polymerase can withstand as it is a DNA polymerase that has been isolated from Thermus aquaticus (hot springs bacteria). • The target area is not completely isolated in the first few cycles of DNA replication and Taq polymerase adds nucleotides until it reaches the end of the DNA.
Advanced Techniques • Variable length strands are produced that start at the target region at one end and extend beyond the target at the other end. • In the second cycle the DNA strands are again heated and separated and primers allowed to anneal. On two of the DNA strands one end terminates at the target region (from cycle one) and the primers anneal to the other end of the target area. • Taq polymerase then adds appropriate nucleotides and ceases when it reaches the end that terminates with the target region. These are constant-length strands. • By the third cycle the number of copies of the targeted strands increases exponentially
Advanced Techniques • Restriction Fragment Length Polymorphism • Polymorphism any difference in DNA sequence, coding or noncoding, that can be detected between individuals • Restriction fragment length polymorphism (RFLP) analysis technique by which DNA regions are digested using restriction enzymes and subjected to radioactive complementary DNA probes to compare the differences in DNA fragment lengths between individuals.
Advanced Techniques • Southern blotting process that allows DNA run on a gel to be transferred to a nylon membrane using electrical current. • This is then placed in a solution containing radioactive complementary nucleotide probes. • Base paring between the DNA and the probes is called hybridization. • After hybridization the blot is exposed to x-ray film and areas that have hybridized will band out…this is called an autoradiogram.
Advanced Techniques • DNA Sequencing • Human Genome ProjectSanger dideoxy method or Chain Termination Technique (1977)…DNA sequencing technique based on DNA replication that uses dideoxy nucleoside triphosphates • DNA template is treated as being single stranded • A short single stranded radioactively labeled primer is added to the end of the DNA template
Advanced Techniques • Identical copies of this complex are placed in four reaction tubes • Each tube has DNA polymerase and a supply of all four deoxynucleoside triphosphates (dATP; dTTP; dGTP & dCTP) • Also, each of the four tubes contains a different radioactively labeled dideoxy analogue (nucleoside triphosphate whose ribose sugar does not possess a hydroxyl group on the 3’ carbon)
Advanced Techniques • What does this do? • Well, if the 3’-OH group is missing (as it is on the ddNTPs) then the phosphodiester bonds can not be continued and the chain will stop growing when one of the ddNTPs are incorporated. Since only a fraction of the dNTPs are dideoxy analogues, different lengths of DNA will be built. Take the following example from the tube that contained ddATP: • 5’ – TTA • TTACGTA • TTACGTACGTA • TTACGTACGTAA – 3’ • These differing lengths of DNA can be separate using gel electrophoresis and analyzed accordingly.
Applications - 6.4 • Medical Applications • Genetic Screening process by which an individual’s DNA is scanned for genetic mutations • Personal information for later in life • Fetal scan • What do you do with this information? • Gene Therapy alteration of a genetic sequence in an organism to prevent or treat a genetic disorder • Chronic Pain pronociceptive transmitters induce pain and antinociceptive transmitters dampen pain • The idea is to introduce antinociceptive transmitters into cells that might cause pain or to areas that produce these transmitters already, but with more genes they will produce more of the transmitter to dampen the pain.
Applications • Agricultural Applications • Nestor and Chilton (1981) created the first transgenic plant • Ti plasmid (tumour inducing) carried by soil bacteria Agrobacterium tumefaciens was used as the vector • Bacteria enter the plant through a wound and create a bulbous growth (crown gall) • Only the T section of the plasmid is incorporated into the plant’s chromosomal DNA however a foreign gene that has been incorporated into the T section therein gets incorporated into the plant’s chromosomal DNA • Only dicotyledon plants (beans, peas, potatoes), monocotyledons (wheat, corn, rice) require a gene gun to incorporate new DNA • Factors now transgenically inserted into plants: hardiness, increased yield, uniformity, insect and virus resistance, herbicide tolerance, antiripening and antiaging
Applications • Forensics • DNA from the crimescene is compared to DNA from potential suspects • DNA fingerprinting pattern of bands on a gel, from RFLP or PCR that is unique for each individual • RFLP requires that the DNA sample is undegraded • PCR requires only minute quantities of DNA that can be degraded • Both utilize the non-coding regions of DNA as they are the locations that the most variability lies. The non-coding regions differ in the quantity of variable number tandem repeats • It is illegal in Canada to refuse to provide a DNA sample to police on arrest if requested