Polymerase Chain Reaction & its applications
650 likes | 1.2k Views
Polymerase Chain Reaction & its applications. Dr.Shubha Gopal Associate Professor & Chairperson Department of Studies in Microbiology University of Mysore Manasagangotri Mysore – 570 006. 15.12.2011 UGC Academic Staff College.

Polymerase Chain Reaction & its applications
E N D
Presentation Transcript
Polymerase Chain Reaction&its applications Dr.Shubha Gopal Associate Professor & Chairperson Department of Studies in Microbiology University of Mysore Manasagangotri Mysore – 570 006 15.12.2011 UGC Academic Staff College
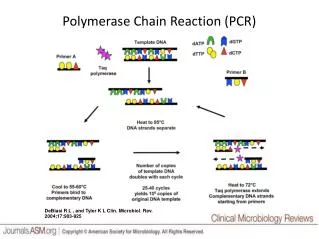
The Polymerase Chain Reaction (PCR) was not a discovery, but rather an invention • A special DNA polymerase (Taq) is used to make many copies of a short length of DNA (100-10,000 bp) defined by primers • Kary Mullis, the inventor of PCR, was awarded the 1993 Nobel Prize in Chemistry
Development…. • PCR work was first published (1985)using Klenow polymerase – unstable with heat • New enzyme had to be added manually at each step • Maximum length 400bp • Great idea – not very practical • First reports using DNA polymerase from Thermus aquaticus (1988) • Taq-polymerase (Saiki et al, 1988) from Yellow stone National Park hot springs • Developed automatic “thermocycler” programmable heat block
Polymerase Chain Reaction (PCR) • PCR is a technique which is used to amplify the number of copies of a specific region of DNA, in order to produce enough DNA to be adequately tested. • The purpose of a PCR is to make a huge number of copies of a gene. As a result, it now becomes possible to analyze and characterize DNA fragments found in minute quantities in places like a drop of blood at a crime scene or a cell from an extinct dinosaur.
What all PCR Can Do ? • Starting with one original copy an almost infinite number of copies can be made using PCR • “Amplified” fragments of DNA can be sequenced, cloned, probed or sized using electrophoresis • Defective genes can be amplified to diagnose any number of illnesses • Genes from pathogens can be amplified to identify them (i.e., HIV, Vibrio sp., Salmonella sp. etc.) • Amplified fragments can act as genetic fingerprints
PCR Reagents • 1X Buffer • 10mM Tris-HCl, 50mM KCl • MgCl2 • 1mM - 4mM (1.5mM) • dNTPs • 200μM • Primers • 100nM-1μM, 200nm (or less) for real time analysis • DNA polymerase • Taq DNA polymerase is thermostable • 1-4 Units (1 unit) • DNA • 10pg-1μg (20ng)
Origin of Replication 5’ 3’ 3’ 5’ 3’ 5’ 5’ 3’ 3’ 5’ 5’ 3’ 3’ 5’ 5’ 3’ 3’ 5’ 3’ 5’ Initiation - Forming the Replication Eye
3’ 5’ 5’ 3’ 3’ 5’ 3’ Primase 5’ Single strand binding proteins Laging Strand 5’ 5’ 3’ 5’ RNA Primers DNA Polymerase 5’ 3’ Helicase Leading Strand 5’ 3’ Extension - The Replication Fork Okazaki fragment
Function PCR How are the functions of replication achieved during PCR ??? . N/A as fragments are short • ENZYMES • Helicase • SSB proteins • Topoisomerase • DNA pol • Primase • Ligase . Heat • Melting DNA • Polymerizing DNA . Taq Polymerase • Providing primer . Primers added to the reaction mix • Joining nicks
Melting 100 94 oC Temperature 50 0 T i m e 3’ 5’ 5’ 3’ PCR
Melting 100 94 oC Temperature 50 0 T i m e 3’ 5’ PCR Heat 5’ 3’
Melting Melting 100 94 oC 94 oC Extension Annealing Primers Temperature 72 oC 50 oC 50 0 T i m e 5’ 3’ 5’ 5’ 5’ 3’ PCR
30x Melting Melting 100 94 oC 94 oC Extension Annealing Primers Temperature 72 oC 50 oC 50 0 T i m e 5’ 3’ 5’ 5’ 5’ 3’ PCR Heat Heat 5’
30x Melting Melting 100 94 oC 94 oC Extension Annealing Primers Temperature 72 oC 50 oC 50 0 T i m e 5’ 3’ 5’ 5’ 5’ 5’ 5’ 5’ 5’ 3’ PCR
30x Melting Melting 100 94 oC 94 oC Extension Annealing Primers Temperature 72 oC 50 oC 50 3’ 5’ 0 5’ T i m e 5’ 5’ 3’ 5’ 5’ 5’ 5’ PCR Heat Heat
30x Melting Melting 100 94 oC 94 oC Extension Annealing Primers Temperature 72 oC 50 oC 50 3’ 5’ 0 5’ T i m e 5’ 5’ 3’ 5’ 5’ 5’ 5’ 5’ 5’ 5’ 5’ PCR
30x Melting Melting 100 94 oC 94 oC Extension Annealing Primers Temperature 72 oC 50 oC 50 3’ 5’ 0 5’ T i m e 5’ 5’ 3’ 5’ 5’ 5’ Fragments of defined length 5’ 5’ 5’ 5’ 5’ PCR
Number of cycles 0 10 15 20 25 30 Size Marker More Cycles = More DNA
PCR Optimisation 1: Buffers • Most buffers have only KCl (50mM) and Tris (10mM) • Concentrations of these can be altered • KCl facilitates primer binding but concentrations higher than 50mM inhibit Taq • DMSO, BSA, gelatin, glycerol, Tween-20, Nonidet P-40, Triton X-100 can be added to aid in the PCR reaction • Enhance specificity, but also can be inhibitory • Pre-mixed buffers are available
PCR Optimisation 2: MgCl2 • MgCl2: required for primer binding • MgCl2 affects primer binding, Tm of template DNA, product- and primer-template associations, product specificity, enzyme activity and fidelity • dNTPs, primers and template chelate and sequester the Mg ion, therefore concentration should be higher than dNTPs (as these are the most concentrated) • Excess magnesium gives non-specific binding • Too little magnesium gives reduced yield
PCR Optimisation 3: Primer Design • Specific to sequence of interest • Length 18-30 nucleotides • Annealing temperature 50oC-70oC • Ideally 58oC-63oC • GC content 40-60% • 3’ end critical (new strand extends from here) • GC clamp (G or C at 3’ terminus) • Inner self complementarity: • Hairpins <5, dimers <9 • 3’ complementarity: • <3-4 bases similar to other primer regions
PCR Optimisation 4: Cycling Conditions • Denaturation: • Some Taq polymerases require initial denaturation (hot start) • Annealing temperature: • ~ 5oC less than Tm of primers • Tm = 4(G + C) + 2(A + T)oC (or use of primer software) • Decrease in annealing temperature result in non-specific binding • Increase in annealing temperature result in reduced yield
PCR Optimisation 5: Cycle Number Theoretical yield = 2n ie. cycle 1 = 2, cycle 2 = 4, cycle 3 = 8, etc eg. if you start with 100 copies after 30 cycles you will have 107, 374, 182, 400 copies • 25-40 cycles • Half-life of Taq is 30 minutes at 95oC • Therefore if you use more than 30 cycles at denaturation times of 1 minute, the Taq will not be very efficient at this point
In summary • Primer length should not exceed 30 mer. • Tm, not more than 60 degree . • GC Content should be in the range of 40-60 % for optimum PCR efficiency. • Primers should end (3′) in a G or C, or CG or GC: this prevents “breathing” of ends and increases efficiency of priming.
GCG PRIME • The GCG program PRIME is a good tool for the design of primers for PCR and sequencing • For PCR primer pair selection, you can choose a target range of the template sequence to be amplified • In selecting appropriate primers, PRIME allows you to specify a variety of constraints on the primer and amplified product sequences. • upper and lower limits for primer and product melting temperatures • primer and product GC contents. • a range of acceptable primer sizes • a range of acceptable product sizes. • required bases at the 3' end of the primer (3' clamp) • maximum difference in melting temperatures between a pair of PCR primers
PC Software • There are a number of (expensive) dedicated PCR primers design programs for personal computers that have “special features” such as nested and multiplex PCR : • Oligo (Molecular Biology Insights, Inc.) • Primer Premier (Premier Biosoft) • Many of the comprehensive MolBio. programs also have PCR features • Mac Vector • OMIGA • Vector NTI • Gene Tool
Primer3 http://frodo.wi.mit.edu/cgi-bin/primer3/primer3_www.cgi
Primer Problems • primers should flank the sequence of interest • primer sequences should be unique • primers that match multiple sequences will give multiple products • repeated sequences can be amplified - but only if unique flanking regions can be found where primers can bind
PCR Based Methods • Sequence Specific Oligonucleotide (SSO) probe • Amplified fragment-length polymorphism to generate finger prints • Large VNTR regions (10-30 b.p. repeat) • Short Tandem Repeats (STR) (2-7 b.p. repeat) • RAPD using universal primers • Rep- PCR (ERIC primers) • PCR- Ribotyping (16S rDNA regions)
Variations of the PCR • Colony PCR • Nested PCR • Multiplex PCR • AFLP PCR • Hot Start PCR • In Situ PCR • Inverse PCR • Asymmetric PCR • Long PCR • Long Accurate PCR • Reverse Transcriptase PCR • Allele specific PCR • Real time PCR
Types of PCR Long PCR: Used to amplify DNA over the entire length up to 25kb of genomic DNA segments cloned. Nested PCR: Involves two consecutive PCR reactions of 25 cycles. The first PCR uses primers external to the sequence of interest. The second PCR uses the product of the first PCR in conjunction with one or more nested primers to amplify the sequence within the region flanked by the initial set of primers. Inverse PCR: Used to amplify DNA of unknown sequence that is adjacent to known DNA sequence. Quantitative PCR: Product amplification w r t time, which is compared with a standard DNA. Hot start PCR: Used to optimize the yield of the desired amplified product in PCR and simultaneously to suppress nonspecific amplification.
Colony PCR Colony PCR- the screening of bacterial (E.Coli) or yeast clones for correct ligation or plasmid products. Pick a bacterial colony with an autoclaved toothpick, swirl it into 25 μl of TE autoclaved dH2O in an microfuge tube. Heat the mix in a boiling water bath (90-100C) for 2 minutes Spin sample for 2 minutes high speed in centrifuge. Transfer 20 μl of the supernatant into a new microfuge tube Take 1-2 μl of the supernatant as template in a 25 μl PCR standard PCR reaction.
Hot Start PCR • This is a technique that reduces non-specific amplification during the initial set up stages of the PCR • The technique may be performed manually by heating the reaction components to the melting temperature (e.g., 95°C) before adding the polymerase • DNA Polymerase- Eubacterial type I DNA polymerase, Pfu • These thermophilic DNA polymerases show a very small polymerase activity at room temperature.
Nested PCR • Two pairs (instead of one pair) of PCR primers are used to amplify a fragment. • First pair -amplify a fragment similar to a standard PCR. Second pair of primers-nested primers (as they lie / are nested within the first fragment) bind inside the first PCR product fragment to allow amplification of a second PCR product which is shorter than the first one. • Advantage- Very low probability of nonspecific amplification
Multiplex PCR • Multiplex PCR is a variant of PCR which enabling simultaneous amplification of many targets of interest in one reaction by using more than one pair of primers.
Inverse PCR • Inverse PCR (Ochman et al., 1988) uses standard PCR (polymerase chain reaction)- primers oriented in the reverse direction of the usual orientation. • The template for the reverse primers is a restriction fragment that has been selfligated • Inverse PCR functions to clone sequences flanking a known sequence. Flanking DNA sequences are digested and then ligated to generate circular DNA. Application • Amplification and identification of flanking sequences such as transposable elements, and the identification of genomic inserts.
Long PCR • Extended or longer than standard PCR, meaning over 5 kilobases (frequently over 10 kb). • Long PCR is useful only if it is accurate. Thus, special mixtures of proficient polymerases along with accurate polymerases such as Pfu are often mixed together. • Application- to clone large genes
Reverse Transcriptase PCR • Based on the process of reverse transcription, which reverse transcribes RNA into DNA and was initially isolated from retroviruses. • First step of RT-PCR - "first strand reaction“-Synthesis of cDNA using oligo dT primers (37°C) 1 hr. • “Second strand reaction“-Digestion of cDNA:RNA hybrid (RNaseH)-Standard PCR with DNA oligo primers. • Allows the detection of even rare or low copy mRNA sequences by amplifying its complementary DNA.
Why real time PCR ? • QUANTITATION OF mRNA • northern blotting • ribonuclease protection assay • in situ hybridization • RT-PCR • most sensitive • can discriminate closely related mRNAs • technically simple • but difficult to get truly quantitative results using conventional PCR