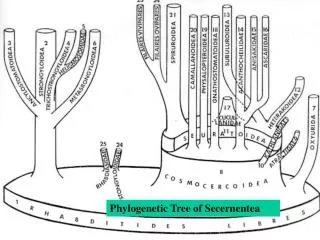
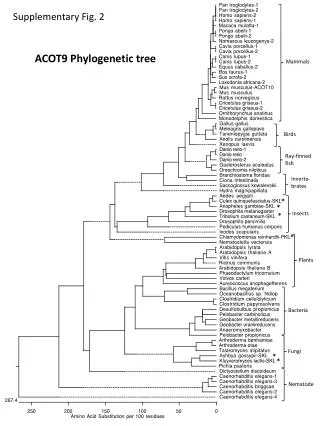
Phylogenetic Tree Construction
Phylogenetic Tree Construction. Dong Xu Computer Science Department 271C Life Sciences Center 1201 East Rollins Road University of Missouri-Columbia Columbia, MO 65211-2060 E-mail: xudong@missouri.edu 573-882-7064 (O) http://digbio.missouri.edu. Outline. Evolution theory

Phylogenetic Tree Construction
E N D
Presentation Transcript
Phylogenetic Tree Construction Dong Xu Computer Science Department 271C Life Sciences Center 1201 East Rollins Road University of Missouri-Columbia Columbia, MO 65211-2060 E-mail: xudong@missouri.edu 573-882-7064 (O) http://digbio.missouri.edu
Outline • Evolution theory • Concept of phylogeny • Molecular clock • Types of trees • UPGMA • Parsimony • Maximum likelihood • An example for bird flu
Evolution • Many theories of evolution • Basic idea: • speciation events lead to creation of different species • Any two species share a (possibly distant) common ancestor
Evolutionary Events • Extinction: A new node u is created at the end of a lineage, no new lineage is started from u • Speciation: A new node u is created at the end of a lineage, and two new lineages are started from u • Hybridization: A new node u is created • when two lineages combine (diploid or polyploid) • when one lineage creates u and the new lineage from u has double the number of homologs (auto-polyploid)
Tree of Life http://tolweb.org/
Toxonomy • Glycine max • Taxonomy ID: 3847Genbank common name: soybeanRank: speciesGenetic code: Translation table 1 (Standard)Mitochondrial genetic code: Translation table 1 (Standard)Other names:common name:soybeans • Lineage( full ) • cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; eudicotyledons; core eudicotyledons; rosids; eurosids I; Fabales; Fabaceae; Papilionoideae; Phaseoleae; Glycine
Kingdom Plantae • Evolutionary tree of plants • From primitive more advanced traits __________ moncot Gymnosperms _______ Non-vascular Dicot Greenalga ancestor Flowers Vascular
Monocot vs. dicot plants (2) • Number of cotyledons: one vs. two
Monocot vs. dicot plants (3) • Leaf venation pattern: • Monocot is parallel • Dicot is net pattern
Monocot vs. dicot plants (4) • Flower parts: • Monocot: in groups of three • Dicot: in groups of four or five
Outline • Evolution theory • Concept of phylogeny • Molecular clock • Types of trees • UPGMA • Parsimony • Maximum likelihood • An example for bird flu
Phylogenies (1) • A phylogeny is a tree that describes the sequence of speciation events that lead to the forming of a set of current day species Aardvark Bison Chimp Dog Elephant
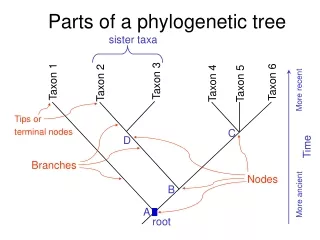
Phylogenies (2) • Leafs - current day species • Nodes - hypothetical most recent common ancestors • Edges length - “time” from one speciation to the next
Tree Terminology d a b c leaf { a,b } edge internal node cluster { a,b,c } root { a,b,c,d }
Rooted/Unrooted Tree • Rooted trees • Single common ancestor • Requires more information • Unrooted trees • Objects are leaves • Internal nodes are some common ancestors • Insufficient information to tell whether not not a given internal node is a common ancestor of any 2 leaves
Motivation • Understand the lineage of different species • Organizing principle to sort species into a taxonomy • Understand how various functions evolved • Understand forces and constraints on evolution • Perform multiple sequence alignment • Predict gene function (phylogenetic footprint)
Tree Basis • Phylogenies are reconstructed based on comparisons between present-day objects • Two main aspects • Topology • How its interior nodes connect to one another and to the leaves • Distance • An estimate of the evolutionary distance between the nodes
Assumptions • homology reflects common ancestry • single common ancestor • treelike relationship exists • positional homology • independent processes • no reversals or convergence • molecular clock
Outline • Evolution theory • Concept of phylogeny • Molecular clock • Types of trees • UPGMA • Parsimony • Maximum likelihood • An example for bird flu
Molecular Clock Theory (1) • For any given protein, accepted mutations in the amino acid sequence for the protein occur at constant rate • Accepted = mutations that allow protein to function without death • Implication # of accepted mutations proportional to length of time interval i.e. relatively constant rateof accepted mutations within a protein
Molecular Clock Theory (2) • Rate of accepted mutations maybe different for different proteins (depending on their tolerance for mutations) • Different parts of a protein may evolve at different rates • Thus, if A and B differ by k accepted mutations, then roughly k/2 mutations have occurred since divergence
Molecular clock Science vol. 289
Outline • Evolution theory • Concept of phylogeny • Molecular clock • Types of trees • UPGMA • Parsimony • Maximum likelihood • An example for bird flu
Species/Gene Trees (1) • Species tree (how are my species related?) • contains only one representative from each species • when did speciation take place? • all nodes indicate speciation events • Gene tree (how are my genes related?) • normally contains a number of genes from a single species • nodes relate either to speciation or gene duplication events
Species/Gene Trees (2) • Your sequence data may not have the same phylogenetic history as the species from which they were isolated • Different genes evolve at different speeds, and there is always the possibility of horizontal gene transfer (hybridization, vector mediated DNA movement, or direct uptake of DNA).
Morphological vs. Molecular • Classical phylogenetic analysis: morphological features • number of legs, lengths of legs, etc. • Modern biological methods allow to use molecular features • Gene sequences • Protein sequences
Dangers in Molecular Phylogenies Gene/protein sequence can be homologous for different reasons: • Orthologs -- sequences diverged after a speciation event • Paralogs -- sequences diverged after a duplication event • Xenologs -- sequences diverged after a horizontal transfer (e.g., by virus)
Ultrametric trees (1) • A metric on a set of objects O given by the assignment of a real number d(x,y) to every pair x,y in O
Ultrametric trees (2) An ultrametric has to fulfill the additional requirement An ultrametric tree is characterized by the three point condition
Additive Trees • Generalization of ultrametric trees • # of mutations were assumed to be proportional to temporal distance of a node to ancestor • Also assumed, mutations took place at same rate in all branches • Additive trees model different rates of mutation along different branches
Additivity • In “real” tree, distances between species are the sum of distances between intermediate nodes k c b j m a i c =
Phylogeny Construction • parsimony methods: fewest changes • likelihood methods: maximize the probability • distance methods: based on pairwise evolutionary distances (sequence similarity, nucleotide composition, etc.)
Outline • Evolution theory • Concept of phylogeny • Molecular clock • Types of trees • UPGMA • Parsimony • Maximum likelihood • An example for bird flu
UPGMA • UPGMA is the unweighted pair group method with arithmetic mean • Distance matrix can come from (e.g) DNA-DNA hybridization, or be constructed from sequence data etc. • Iteratively group the most closely related groups. The average distance between elements in two groups is the distance between the groups.
UPGMA Procedure • find closest pair of units (species, to start with) • connect this pair, defining an evolutionary unit (branch) • compute distances from the ancestor of this unit to all other ungrouped units --Branch length is distance/2 • go back to #1 and repeat
Evolutionary distances among primates (1) nucleotide substitutions per 100 sites H C Humans and chimps are closest: lump them and recompute distances
Evolutionary distances among primates (2) • e.g., (H-C) to gorilla distance • = (H-G+C-G)/2 • = (1.51+1.57)/2 = 1.54 • Gorilla is closest to H-C clade • (((H, C), 1.45), G, 1.54) G H C
Evolutionary distances among primates (3) R O G H C Human-Chimp-Gorilla is closer to Orang than to Rhesus
UPGMA Clustering • Let Ci and Cj be clusters, define distance between them to be • When we combine two cluster, Ci and Cj, to form a new cluster Ck, then
UPGMA: conclusions • UPGMA gives branch lengths or evolutionary distances as well as branching order • if(a big if) mutations occur at a constant rate, we can estimate dates of divergence from sequence differences
Outline • Evolution theory • Concept of phylogeny • Molecular clock • Types of trees • UPGMA • Parsimony • Maximum likelihood • An example for bird flu
Possible Evolutionary Tree (1) t1 t2 t1 t1 t3 t2 t4 t4 t4 t3 t2 t3 t1 1 three-taxa tree t2 1*(2*3-3) = 3 four-taxa trees t3
Possible Evolutionary Tree (3) Taxa (n): 4 2 3 Taxa (n) Unrooted/rooted 2 1/1 3 1/3 4 3/15
Maximum parsimony (1) • Minimizes the number of steps required to generate the observed variation in the sequences • Guaranteed to find the "best" tree - danger of over-fitting the data • Columns representing greater variation dominate • Works best for small, highly conserved sequences
Maximum parsimony (2) • Begin with a multiple sequence alignment • Identify informative sites within the sequences • Tree requiring smallest number of changes identified • Repeat over all informative sites • Length = sum of the # of steps in each branch • Choose tree with smallest length