V8: Virus Structure and Assembly
310 likes | 485 Views
V8: Virus Structure and Assembly. At the simplest level, the function of the outer shells ( CAPSID ) of a virus particle is to protect the fragile nucleic acid genome from: Physical damage - Shearing by mechanical forces.

V8: Virus Structure and Assembly
E N D
Presentation Transcript
V8: Virus Structure and Assembly At the simplest level, the function of the outer shells (CAPSID) of a virus particle is to protect the fragile nucleic acid genome from: Physical damage - Shearing by mechanical forces. Chemical damage- UV irradiation (from sunlight) leading to chemical modification. Enzymatic damage - Nucleases derived from dead or leaky cells or deliberately secreted by vertebrates as defence against infection. The protein subunits in a virus capsid are multiply redundant, i.e. present in many copies per particle. Damage to one or more subunits may render that particular subunit non-functional, but does not destroy the infectivity of the whole particle. Furthermore, the outer surface of the virus is responsible for recognition of the host cell. Initially, this takes the form of binding of a specific virus-attachment protein to a cellular receptor molecule. However, the capsid also has a role to play in initiating infection by delivering the genome from its protective shell in a form in which it can interact with the host cell. http://www-micro.msb.le.ac.uk/109/structure.html Optimization, Energy Landscapes, Protein Folding
Virus Design To form an infectious particle, a virus must overcome two fundamental problems: (1) To assemble the particle utilizing only the information available from the components which make up the particle itself (capsid + genome). (2) Virus particles form regular geometric shapes, even though the proteins from which they are made are irregularly shaped. How do these simple organisms solve these difficulties? The information to answer these problem lie in the rules of symmetry. In 1957, Fraenkel-Conrat & Williams showed that when mixtures of purified tobacco mosaic virus (TMV) RNA & coat protein were incubated together, virus particles formed. The discovery that virus particles could form spontaneously from purified subunits without any extraneous information indicated that the particle was in the free energy minimum state & was therefore the favoured structure of the components. This stability is an important feature of the virus particle. Although some viruses are very fragile & are essentially unable to survive outside the protected host cell environment, many are able to persist for long periods, in some cases for years. http://www-micro.msb.le.ac.uk/109/structure.html Optimization, Energy Landscapes, Protein Folding
Helical capsids Tobacco mosaic virus (TMV) is representative of one of the two major structural classes seen in viruses of all types, those with helical symmetry. The simplest way to arrange multiple, identical protein subunits is to use rotational symmetry & to arrange the irregularly shaped proteins around the circumference of a circle to form a disc. Multiple discs can then be stacked on top of one another to form a cylinder, with the virus genome coated by the protein shell or contained in the hollow centre of the cylinder. Closer examination of the TMV particle by X-ray crystallography reveals that the structure of the capsid actually consists of a helix rather than a pile of stacked disks. A helix can be defined mathematically by two parameters: amplitude (diameter) & pitch (the distance covered by each complete turn of the helix) http://www-micro.msb.le.ac.uk/109/structure.html Optimization, Energy Landscapes, Protein Folding
Helical capsids Helices are rather simple structures formed by stacking repeated components with a constant relationship (amplitude & pitch) to one another - note that if this simple constraint is broken a spiral forms rather than a helix - unsuitable for containing a virus genome. The fact that helical symmetry is a useful way of arranging a single protein subunit to form a particle is confirmed by the large number of different types of virus which have evolved with this capsid arrangement. TMV particles are rigid, rod-like structures, but some helical viruses demonstrate considerable flexibility & longer helical virus particles are often seen to be curved or bent. Flexibility is important attribute since long helical particles are subject to damage from shear forces & the ability to bend reduces the chance of breakage. http://www-micro.msb.le.ac.uk/109/structure.html Optimization, Energy Landscapes, Protein Folding
Icosahedral (isometric) capsids An alternative way of building a virus capsid is to arrange protein subunits in the form of a hollow quasi-spherical structure, enclosing the genome within. The criteria for arranging subunits on the surface of a solid are more complex than those for building a helix. In the 1950s, Brenner & Horne developed sophisticated techniques which enabled them to use electron microscopy to reveal many of the fine details of the structure of virus particles. One of the most useful techniques proved to be the use of electron-dense dyes such as phosphotungstic acid or uranyl acetate to examine virus particles by negative staining. The small metal ions in such dyes are able to penetrate the minute crevices between the protein subunits in a viral capsid to reveal the fine structure of the particle. http://www-micro.msb.le.ac.uk/109/structure.html Optimization, Energy Landscapes, Protein Folding
Icosahedral viruses Francis Crick & James Watson (1956), were the first to suggest that virus capsids are composed of numerous identical protein sub-units arranged either in helical or cubic (=icosahedral) symmetry. In order to construct a capsid from repeated subunits, a virus must 'know the rules' which dictate how these are arranged. For an icosahedron, the rules are based on the rotational symmetry of the solid, which is known as 2-3-5 symmetry: An axis of two-fold rotational symmetry through the centre of each edge An axis of three-fold rotational symmetry through the centre of each face An axis of five-fold rotational symmetry through the centre of each corner (vertex) http://www-micro.msb.le.ac.uk/109/structure.html Optimization, Energy Landscapes, Protein Folding
Icosahedral viruses The simplest icosahedral capsids are built up by using 3 identical subunits to form each triangular face, thereby requiring 60 identical subunits to form a complete capsid. A few simple virus particles are constructed in this way, e.g. bacteriophage ØX174: http://www-micro.msb.le.ac.uk/109/structure.html Optimization, Energy Landscapes, Protein Folding
Examples of icosahedral viruses In most cases, analysis reveals that icosahedral virus capsids contain more than 60 subunits, for the reasons of genetic economy given above. The capsids of picornaviruses provide a good illustration of the construction of icosahedral virus particles (e.g. polioviruses, foot-and-mouth disease virus, rhinoviruses). http://www-micro.msb.le.ac.uk/109/structure.html Optimization, Energy Landscapes, Protein Folding
Enveloped viruses 'Naked' virus particles, i.e. those in which the capsid proteins are exposed to the external environment are produced from infected cells at the end of the replicative cycle when the cell dies, breaks down & lyses, releasing the virions which have been built up internally. This simple strategy has drawbacks. In some circumstances it is wasteful, resulting in the premature death of the cell. http://www-micro.msb.le.ac.uk/109/structure.html Optimization, Energy Landscapes, Protein Folding
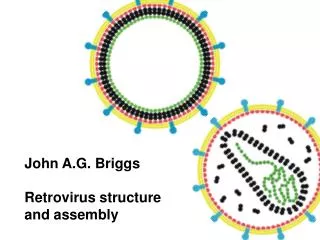
Enveloped viruses Many viruses have devised strategies to exit the infected cell without its total destruction. This presents a difficulty in that all living cells are covered by a membrane composed of a lipid bilayer. The viability of the cell depends on the integrity of this membrane. Viruses leaving the cell must therefore allow this membrane to remain intact & this is achieved by extrusion (budding) of the particle through the membrane, during which process the particle becomes coated in a lipid envelope derived from the host cell membrane & with a similar composition: http://www-micro.msb.le.ac.uk/109/structure.html Optimization, Energy Landscapes, Protein Folding
Enveloped viruses The structure underlying the envelope may be based on helical or icosahedral symmetry & may be formed before or as the virus leaves the cell. In the majority of cases, enveloped viruses use cellular membranes as sites allowing them to direct assembly. The formation of the particle inside the cell, maturation & release are in many cases a continuous process. The site of assembly varies for different viruses. Not all enveloped viruses bud from the cell surface membrane, many viruses use cytoplasmic membranes such as the golgi complex, others such as herpesviruses which replicate in the nucleus may utilize the nuclear membrane. In these cases, the virus is usually extruded into some form of vacuole, in which it is transported to the cell surface & subsequently released. http://www-micro.msb.le.ac.uk/109/structure.html Optimization, Energy Landscapes, Protein Folding
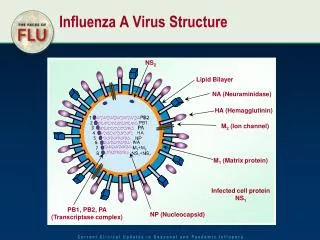
Envelope Proteins Envelope: If the virus particle became covered in a smooth, unbroken lipid bilayer, this would be its undoing. Such a coating is effectively inert, & though effective as a protective layer preventing desiccation of or enzymatic damage to the particle, would not permit recognition of receptor molecules on the host cell. Therefore, viruses modify their lipid envelopes by the synthesis of several classes of proteins which are associated in one of three ways with the envelope: Matrix Proteins: These are internal virion proteins whose function is effectively to link the internal nucleocapsid assembly Glycoproteins: These are transmembrane proteins, anchored to the membrane by a hydrophobic domain & can be subdivided into two types, by their function: External Glycoproteins - Anchored in the envelope by a single transmembrane domain. Most of the structure of the protein is on the outside of the membrane, with a relatively short internal tail. Often individual monomers associate to form the 'spikes' visible on the surface of many enveloped viruses in the electron microscope. Such proteins are the major antigens of enveloped viruses. Transport Channels - This class of proteins contains multiple hydrophobic transmembrane domains, forming a protein-lined channel through the envelope, which enables the virus to alter the permeability of the membrane, e.g. ion-channels. http://www-micro.msb.le.ac.uk/109/structure.html Optimization, Energy Landscapes, Protein Folding
Complex virus structures • The majority of viruses can be fitted into one of the three structural classes: • helical symmetry, • icosahedral symmetry or • enveloped viruses based on either of these two. • However, there are many viruses whose structure is more complex. • In these cases, although the general principles of symmetry already described are often used to build part of the virus shell, the larger & more complex viruses cannot be simply defined by a mathematical equation as can a simple helix or icosahedron. • Because of the complexity of some of these viruses, they have defied attempts to determine detailed atomic structures using the techniques described earlier. http://www-micro.msb.le.ac.uk/109/structure.html Optimization, Energy Landscapes, Protein Folding
Pox viruses An example of such a group & the problems of complexity is shown by the members of the poxvirus family. These viruses have oval or 'brick-shaped' particles 200 - 400 nm long. In fact, these particles are so large that they were first observed using high resolution optical microscopes in 1886 & thought at that time to be 'the spores of micrococci'. http://www-micro.msb.le.ac.uk/109/structure.html Optimization, Energy Landscapes, Protein Folding
Pox viruses The external surface of the virion is ridged in parallel rows, sometimes arranged helically. The particles are extremely complex & have been shown to contain more than 100 different proteins. Antigenically, poxviruses are very complex, inducing both specific & cross-reacting antibodies - hence ability to vaccinate against one disease with another virus (i.e. the use of vaccinia virus to immunize against smallpox (variola) virus). Poxviruses & a number of other complex viruses also emphasise the true complexity of some virus - there are at least ten enzymes present in poxvirus particles, mostly concerned with nucleic acid metabolism/genome replication. http://www-micro.msb.le.ac.uk/109/structure.html Optimization, Energy Landscapes, Protein Folding
Summary 1. To protect the genome 2. To deliver the genome to the appropriate site in the host cell so that it can be replicated. 3. A number of repeated structural motifs found in many different virus groups are evident. The most obvious is the division of many virus structures into those based on helical or icosahedral symmetry. 4. Virus particles are not inert. Many are armed with a variety of enzymes which carry out a range of complex reactions, most frequently concerned with the replication of the genome. http://www-micro.msb.le.ac.uk/109/structure.html Optimization, Energy Landscapes, Protein Folding
Icosahedral Viruses • Half of all virus families share icosahedral geometry, even though they may have nothing else in common • speculation that there may be a physical advantage to icosahedral geometries. Why is it interesting to understand these laws? • One may be able to develop strategies to fight viruses. • One could build self-assembling nano-particles from proteins with special desired properties. http://www-micro.msb.le.ac.uk/109/structure.html Optimization, Energy Landscapes, Protein Folding
Icosahedral Geometry An icosahedron is 20-sided solid, where each facet has threefold symmetry (Fig. 1A). To build an icosahedron out of protein, each face must be made of at least three proteins, because an individual protein cannot have intrinsic threefold symmetry. In an icosahedron, the proteins are related by exact two-, three-, and fivefold symmetry axes. It turns out that few spherical viruses are built of 60 subunits, but most viruses are built of T multiples of 60, where the T (for triangulation) number indicates the number of subunits within each of the 60 icosahedral asymmetric units. The term quasi-equivalence indicates that the subunits are in distinct but quasi-equivalent environments. In this manner, some subunits are arranged around icosahedral fivefold axes and others are arranged as hexamers. http://www-micro.msb.le.ac.uk/109/structure.html Optimization, Energy Landscapes, Protein Folding
Icosahedral Geometry Quasi-equivalence is readily apparent in a virus structure by the presence of pentameric and hexameric groupings of subunits, the capsomers (Fig. 1B). A selection of cryoelectron microscopy image reconstructions demonstrating different T numbers: polio (T=1), small hepatitis B virus (HBV) capsid (T=3), large HBV capsid (T=4), bacteriophage HK97 (T=7), and herpes simplex virus (T=16). An icosahedral facet is highlighted on selected images. http://www-micro.msb.le.ac.uk/109/structure.html Optimization, Energy Landscapes, Protein Folding
Icosahedral Symmetry of a viral capsid (a) Cryo-TEM reconstruction of CCMV. (b) Arrangement of subunits on a truncated icosahedron; A, B, and C denote the three symmetry nonequivalent sites. Zandi et al., PNAS 101, 15556 (2004) Optimization, Energy Landscapes, Protein Folding
Why do viruses adopt icosahedral symmetry? Derive model for equilibrium viral structures that retains the essential features of the process and results in the predominance of icosahedral CK structures as well as the existence of other structures observed in vitro that do not fall into this classification. Start from the observation that, despite the wide range of amino acid sequences and folding structures of viral coat proteins, capsid proteins spontaneously self-assemble into a common viral architecture. The actual kinetic pathways and intermediates involved are quite varied [e.g., CCMV assembles from dimers, Polyoma from pentameric capsomers, and HK97 from pentamers and hexamers] but the equilibrium structures of viral capsids are invariably made up of the same units (e.g., pentamers andor hexamers). This finding suggests that, although the interaction potential between subunits is asymmetric and species-specific, capsomers interact through a more isotropic and generic interaction potential. The focus of the present work is not on the kinetics process of the assembly but rather on understanding the optimal equilibrium structures. Zandi et al., PNAS 101, 15556 (2004) Optimization, Energy Landscapes, Protein Folding
Monte Carlo simulations Based on the ideas noted above we consider a minimal model for the equilibrium structure of molecular shells in which we do not attempt to describe individual subunits but instead focus on the capsomers. The effective capsomer-capsomer potential V(r) is assumed to depend only on the separation r between the capsomer centers and captures the essential ingredients of their interaction: a short-range repulsion, representing subunit conformational rigidity, plus a longer-range attraction, corresponding to the driving force (e.g., hydrophobic interaction) for capsomer aggregation. The capsomer-capsomer binding energy 0 is taken to be 15 kT, a typical value reported from atomistic calculations of subunit binding energies. Zandi et al., PNAS 101, 15556 (2004) Optimization, Energy Landscapes, Protein Folding
Two different morphological units Another essential feature of viral capsids is the existence of two different morphological units (pentamers and hexamers). To account for the intrinsic differences between capsomer units we assume that they can adopt two internal states: P(entamer) and H(examer). The potential has the same form for interactions between different capsomer types except that the equilibrium spacing [the minimum of V(r)] includes the geometrical size difference between pentamers and hexamers of the same edge length (size ratio 0.85). The energy difference E between a P and an H capsomer, which reflects differences between individual contact interactions and folding conformations of pentamer and hexamer proteins, enters as a Boltzmann factor eE/kTthat provides the relative thermal probability for a noninteracting unit to be in the P state. For each fixed total number of capsomers N, the number NP of P units (and hence the number NH = N -NP of H units) is permitted to vary and was not fixed to be 12 (as in the CK construction). Zandi et al., PNAS 101, 15556 (2004) Optimization, Energy Landscapes, Protein Folding
Monte Carlo simulations We carry out Monte Carlo simulations in which N interacting capsomers are allowed to range over a spherical surface while switching between P and H states, thus exploring all possible geometries and conformations. In this way we obtain the optimal structure for a given number N of capsomers and a given capsid radius R. We have used Metropolis Monte Carlo (MC) simulation with 105 equilibration steps and 105 production steps. An elementary MC step consisted of either an attempt to move a randomly chosen disk over the surface of a sphere in a random direction or an attempt to change its size. The ratio of MC attempts of moving a disk versus switching the size of a disk was set to 10. However, we tested different ratios and the result was robust, independent of the ratio. The finite-temperature internal energy E(R) is evaluated for each of a range of equilibrated sphere radii R and then minimized with respect to R, leading to a special radius R* for each N. We tested different forms for V(r) and found the conclusions discussed below to be robust. Zandi et al., PNAS 101, 15556 (2004) Optimization, Energy Landscapes, Protein Folding
Monte Carlo simulations Note favorable numbers of particles. Energy per capsomer for E =0 (black curve) and |E / o | large compared to one (dotted curve). Zandi et al., PNAS 101, 15556 (2004) Optimization, Energy Landscapes, Protein Folding
Icosahedral Symmetry of a viral capsid Minimum energy structures produced by Monte Carlo simulation, with P-state capsomers shown in black. (a) The P and H states here have the same energies. The resulting N=12, 32, 42, and 72 structures correspond to T=1, T=3, T=4, and T=7 C-K icosahedra. (b) Minimum energy structures for |E / o | >> 1, i.e., only one size of capsomer. The N = 24 and 48 structures have octahedral symmetry, and N = 32 is icosahedral, whereas N = 72 is highly degenerate, fluctuating over structures with different symmetry, including T = 7. Zandi et al., PNAS 101, 15556 (2004) Optimization, Energy Landscapes, Protein Folding
Icosahedral Symmetry of a viral capsid Stability around the minimum 72. N = 71 has essentially the same structure as N = 72, but a structural defect. N = 73 has many surface defects. Zandi et al., PNAS 101, 15556 (2004) Optimization, Energy Landscapes, Protein Folding
Mechanical properties of capsids Our minimal model for capsid structure posits an explicit potential for capsomer interactions, which provides us with a tool to study the mechanical and genome release properties of viral capsids. To address the effect of strain on capsid structure, we repeated our E0 simulations at each of successive fixed capsid radii in excess of the optimal radius R*. For fixed N 32 (T = 3) and NP = 12, the capsid bursts dramatically when RR* exceeds a critical value (1.107), in the form of a large crack stretching across the capsid surface. The bursting of the capsid is one of several possible gene release scenarios. Just before this point is reached the capsid is uniformly swollen (see Fig. 6a), with all interstitial holes grown larger in size compared to those in the optimized R R* structure (see N = 32 in Fig. 3a). The appearance of these pores constitutes another mechanism for genome release. Zandi et al., PNAS 101, 15556 (2004) Optimization, Energy Landscapes, Protein Folding
Mechanical properties of capsids Finally, when we allow the number of capsomers to change during swelling, we find that the bursting scenario competes with still another mechanism, decapsidation; at a critical radius 1.107 R* the capsid energy can be reduced by ejecting one of the 12 pentamers, followed by a decrease in capsid size. These phenomena have been observed, for instance, for the Tymoviruses and a series of Flock House virus mutants. The fact that the minimal model reproduces realistic release mechanisms, in addition to accounting for both the predominant T-structures and the exceptional nonicosahedral structures, suggests that it can be applied as well to studying the mechanical properties of capsids and serve as a guide for the design of artificial viruses. Zandi et al., PNAS 101, 15556 (2004) Optimization, Energy Landscapes, Protein Folding