100
160 likes | 396 Views
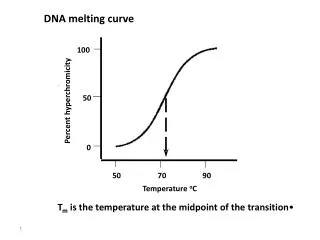
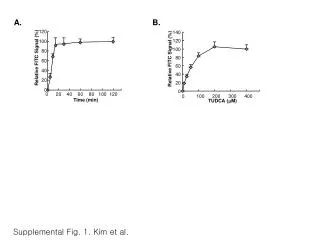
DNA melting curve. 100. 50. Percent hyperchromicity. 0. 50. 70. 90. Temperature o C. T m is the temperature at the midpoint of the transition. T m is dependent on the G-C content of the DNA. E. coli DNA is 50% G-C. Percent hyperchromicity. 50. 60. 70. 80.

100
E N D
Presentation Transcript
DNA melting curve 100 50 Percent hyperchromicity 0 50 70 90 Temperature oC • Tm is the temperature at the midpoint of the transition
Tm is dependent on the G-C content of the DNA E. coli DNA is 50% G-C Percent hyperchromicity 50 60 70 80 Temperature oC Average base composition (G-C content) can be determined from the melting temperature of DNA
Genomic DNA, Genes, Chromatin a). Complexity of chromosomal DNA i). DNA reassociation ii).Repetitive DNA and Alu sequences iii). Genome size and complexity of genomic DNA b). Gene structure i). Introns and exons ii). Properties of the human genome iii). Mutations caused by Alu sequences c). Chromosome structure - packaging of genomic DNA i). Nucleosomes ii). Histones iii). Nucleofilament structure iv). Telomeres, aging, and cancer
Complexity of chromosomal DNA:DNA reassociation (renaturation) • The complexity of a DNA is a function of how many base pairs it has. • In low complexity = few hundred • High complexity = millions of base pairs. • Low complexity sequences are able to find each other much faster during a renaturation reaction, than are high complexity sequences, since the low complexity sequences have to make fewer collisions to find a base-pairing partner. • Thus, the rate of a renaturation reaction is a function of the complexity of the DNA. Or in other words, the complexity of a DNA can be determined by measuring its second-order reassociation rate constant (k2).
DNA reassociation (renaturation) Double-stranded DNA Denatured, single-stranded DNA Faster, zippering reaction to form long molecules of double- stranded DNA k2 Slower, rate-limiting, second-order process of finding complementary sequences to nucleate base-pairing
Type of DNA % of Genome Features 1. Single-copy (unique) ~75% Includes most genes 1 2. Repetitive a- Interspersed ~15% Interspersed throughout genome between and within genes; includes Alusequences 2and VNTRs or mini (micro) satellites b- Satellite (tandem) ~10% Highly repeated, low complexity sequences usually located in centromeres and telomeres 2Alu sequences are about 300 bp in length and are repeated about 300,000 times in the genome. They can be found adjacent to or within genes in introns or nontranslated regions. 1 Some genes are repeated a few times to thousands-fold and thus would be in the repetitive DNA fraction 0 fast ~10% intermediate ~15% 50 slow (single-copy) ~75% 100 I IIIIIIII
Classes of repetitive DNA • Interspersed repeats are sequences that are repeated many times and scattered throughout the genome. • In contrast, tandem repeats are sequences that are repeated many times adjacent to each other. They are usually found in the centromeres and telomeres of chromosomes (the sequence above TTAGGG comprises human teleomeric DNA
Classes of repetitive DNA Interspersed (dispersed) repeats (e.g., Alu sequences) GCTGAGG GCTGAGG GCTGAGG Tandem repeats (e.g., microsatellites) TTAGGGTTAGGGTTAGGGTTAGGG
Human Genome Project • Knowing the complete sequence of the human genome will: • allow medical researchers to more easily find disease-causing genes. • understand how differences in our DNA sequences from individual to individual may affect our predisposition to diseases and our ability to metabolize drugs. • Because the human genome has ~3 billion bp of DNA and there are 23 pairs of chromosomes in diploid human cells, the average metaphase chromosome has ~130 million bp DNA.
Genome sizes in nucleotide pairs (base-pairs) plasmids viruses bacteria fungi plants algae insects mollusks bony fish The size of the human genome is ~ 3 X 109bp; almost all of its complexity is in single-copy DNA. The human genome is thought to contain ~30,000 to 40,000 genes. amphibians reptiles birds mammals 104 105 106 107 108 109 1010 1011
Gene Structure • Most genes in the human genome are called "split genes" because they are composed of "exons" separated by "introns." • The exons are the regions of genes that encode information that ends up in mRNA. • The transcribed region of a gene (double-ended arrow) starts at the +1 nucleotide at the 5' end of the first exon and includes all of the exons and introns (initiation of transcription is regulated by the promoter region of a gene, which is upstream of the +1 site). • RNA processing (the subject of a another lecture) then removes the intron sequences, "splicing" together the exon sequences to produce the mature mRNA. • The translated region of the mRNA (the region that encodes the protein) is indicated in blue. Note that there are untranslated regions at the 5' and 3‘ ends of mRNAs that are encoded by exon sequence but are not directly translated.
Gene structure promoter region exons (filled and unfilled boxed regions) +1 introns (between exons) transcribed region mRNA structure 5’ 3’ translated region
The (exon-intron-exon)n structure of various genesintrons can be very long, while exons are usually relatively short. • Next figure shows examples of the wide variety of gene structures seen in the human genome. Some (very few) genes do not have introns. One example is the histone genes, which encode the small DNA-binding proteins, histones H1, H2A, H2B, H3, and H4. • Shown here is a histone gene that is only 400 base pairs (bp) in length and is composed of only one exon. • The beta-globin gene has three exons and two introns. • The hypoxanthine-guanine phosphoribosyltransferase (HGPRT or HPRT) gene has nine exons and is over 100-times larger than the histone gene, yet has an mRNA that is only about 3-times larger than the histone mRNA (total exon length is 1,263 bp). This is due to the fact that introns can be very long, while exons are usually relatively short. • An extreme example of this is the factor VIII gene which has numerous exons (the blue boxes and blue vertical lines).
The (exon-intron-exon)n structure of various genes introns can be very long, while exons are usually relatively short. histone total = 400 bp; exon = 400 bp b-globin total = 1,660 bp; exons = 990 bp HGPRT (HPRT) total = 42,830 bp; exons = 1263 bp factor VIII total = ~186,000 bp; exons = ~9,000 bp
Properties of the human genome • Nuclear genome • the haploid human genome has ~3 X 109bp of DNA • single-copy DNA comprises ~75% of the human genome • the human genome contains ~20,000 to 25,000 genes • most genes are single-copy in the haploid genome • genes are composed of from 1 to >75 exons • genes vary in length from <100 to >2,300,000 bp • Alu sequences are present throughout the genome • Mitochondrial genome • circular genome of ~17,000 bp • contains <40 genes