Material and Methods
PATTERN RECOGNITION METHODS FOR THE IDENTIFICATION OF MARINE BIVALVE LARVAE *J. Campbell 1 , J. Slater 2 , J. Gillespie 2 , I.F. Bendezu 2 and F. Murtagh 3 1 Department of Computing, Letterkenny Institute of Technology, Port Road, Co. Donegal, Ireland.

Material and Methods
E N D
Presentation Transcript
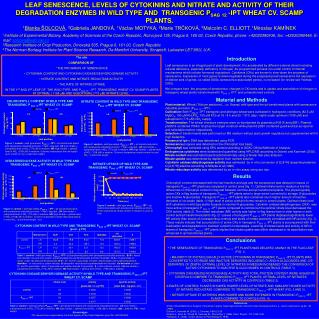
PATTERN RECOGNITION METHODS FOR THE IDENTIFICATION OF MARINE BIVALVE LARVAE *J. Campbell1, J. Slater2, J. Gillespie2, I.F. Bendezu2 and F. Murtagh3 1Department of Computing, Letterkenny Institute of Technology, Port Road, Co. Donegal, Ireland. 2Department of Science, Letterkenny Institute of Technology, Port Road, Co. Donegal, Ireland. 3School of Computer Science, Queen's University Belfast, Belfast, BT7 1NN * Email: jonathan.campbell@lyit.ie Abstract Morphological identification of bivalve larvae to the species level is difficult because of their size, similar shape, similar colour and because of the absence of protruding parts to aid in the identification. Consequently the process is both time-consuming and requires the availability of personnel with considerable experience in the field. The application of computational pattern recognition methods to the automated identification and size analysis of scallop larvae was investigated. For identification, the shape features used were binary invariant moments i.e. the features are invariant to shift (position within the image), scale (induced either by growth or differential image magnification) and rotation. Images of scallop and non-scallop larvae covering a range of sizes were analysed. Scatter plots of selected features indicated the potential for automatic identification. Introduction Aquaculture production is expanding rapidly as the way forward for the marine food sector to develop a sustainable industry for present and future generations. Production valued in excess of €114 million occurred in 2002 and is projected to increase to 97,023 tonnes valued at over €175 million by 2008. Approximately 60% of aquaculture volume comprises shellfish with much of this production, dependent on the collection of wild seed as a source of raw material. To predict when, where and in what quantities wild shellfish seed will be available requires knowledge of the spawning time of the adults and the dispersal patterns of planktonic bivalve larvae in the waters around our coast. Whilst large numbers of plankton samples are relatively easy to collect and good taxonomic keys exist for many of the specimens in such samples, one of the most difficult groups to identify, particularly at the species level, are the Bivalvia. This difficulty arises from the fact that fundamentally all bivalve larvae have a similar shape, they are all brownish in colour to one degree or another, and there are no protruding parts or appendages, which can be used to aid identification (Fig. 1). Morphological methods of identification using a light microscope have been developed for some commercial species but their widespread application is limited by the expertise required and the time-consuming nature of the process. The practical application of methods involving scanning electron microscopy are restricted by the time required to process individual larvae. Molecular methods, both immunological and DNA-based methods, are under development. Fig. 1. Bivalve larvae of a range of species in a natural plankton sample and including amongst others both scallop and mussel larvae. Material and Methods Plankton samples were collected in the North Water of Mulroy Bay, Co. Donegal. A sample consisted of larvae from three vertical hauls, from seabed to water surface (0-22m), collected using a 63um plankton net. Bivalve larvae were preserved with formalin, concentrated by gentle stirring, removed to a 30ml container, examined and photographed using an Olympus BX 51 microscope equipped with digital camera. Raw images of individual bivalve larvae were segmented i.e. separated from their background, using a simple threshold technique based on the image histogram. This reduced each image to a planar, two-dimensional shape from which shape features could be extracted. Shift- and rotation-invariant features were required since larvae can occur at any orientation. Invariant moments of a binarised object, which ignore texture, were used since visual inspection suggests that discrimination should be possible based on outline shape. Following feature extraction, feature vectors were analysed bythestatistical software package R. Fig. 2. Raw Images Fig. 3. Segmented Images Results Normalised invariant moment features from twenty bivalve larval images, five of them scallop larvae, are provided (Fig. 4). A scatter plot of invariant moments h1 and h4 shows the scallop larval data to be tightly clustered and separate from the other data (Fig. 5). Data analysis Shift invariance was achieved by referencing all moment calculations to the centre of mass to yield normalised centralised moments. Scale-invariance was achieved by normalising with respect to area. Rotationally-invariant moments were derived from the normalised central moments. Finally, invariant moments were subjected to normalisation transformations to obtain values of broadly comparable size and range. Discussion Assuming that this small data set is representative, a classification rule to identify scallop larvae could be prepared. Once distinguished from other larvae, the use of area or some other size-related parameter could be used to determine the stage of development and consequently predict the scallop settlement time. Further work on the project will focus on enlargement of the data set, the implementation and evaluation of classification techniques, and the development of a robust segmentation technique based on Markov random field models which combine spatial contiguity with colour/grey level similarity to overcome difficulties where some parts of the object are less optically dense than the background. Fig. 4. Table of Invariant Moment Features. Scallop larvae are highlighted in red. Fig. 5. Scatter Plot -Invariant Moment h1 vs. h4 Scallop larvae are highlighted in red. Conclusions Microscopic images of bivalve larvae of a range of sizes have been analysed using invariant moment features. Based on a limited data set, the distribution of selected features suggests that automated identification of scallop larvae from non-scallop larvae may be possible. Acknowledgments: Financial support for the morphological identification and photomicroscopy of bivalve larvae was provided by the H.E.A. Technological Sector Research Programme Strand III : Core Research Strengths Enhancement supported by the National Development Plan 2000-2006.