0.05
0.05. Sample Collection. Sample processing. SNP association via Taqman assay ( ASPM and HAR1). Sequencing (HAR1). Genes showing accelerated evolutionary properties associate with language.

0.05
E N D
Presentation Transcript
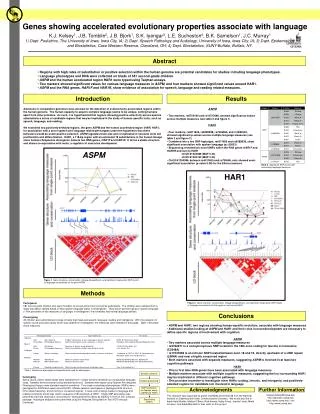
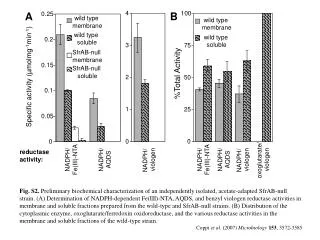
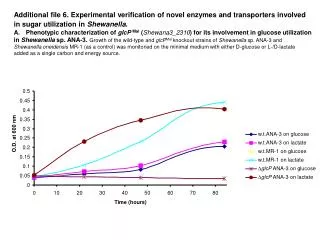
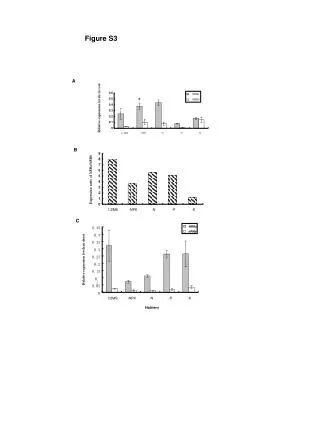
0.05 Sample Collection Sample processing SNP association via Taqman assay (ASPM and HAR1) Sequencing (HAR1) Genes showing accelerated evolutionary properties associate with language K.J. Kelsey1, J.B. Tomblin2, J.B. Bjork1, S.K. Iyengar3, L.E. Sucheston4, B.K. Samelson1, J.C. Murray1 1) Dept. Pediatrics, The University of Iowa, Iowa City, IA; 2) Dept. Speech Pathology and Audiology, University of Iowa, Iowa City, IA; 3) Dept. Epidemiology and Biostatistics, Case Western Reserve, Cleveland, OH; 4) Dept. Biostatistics, SUNY-Buffalo, Buffalo, NY. Abstract • Regions with high rates of substitution or positive selection within the human genome are potential candidates for studies including language phenotypes. • Language phenotypes and DNA were collected on triads of 581 second-grade children. • ASPM and the human accelerated region HAR1 were typed using Taqman assays. • Two markers showed significant values for various language measures in ASPM, and four markers showed significant values around HAR1. • ASPM and the RNA genes, HAR1F and HAR1R, show evidence of association for speech, language and reading related measures. Introduction Results Advances in comparative genomics have allowed for the detection of evolutionarily-accelerated regions within the human genome. The human capacity to acquire complex language seems to be unique, setting humans apart from other primates. As such, it is hypothesized that regions showing positive-selectivity across species substantiate a series of candidate regions that may be implicated in the study of human-specific traits, such as speech, language, and reading. We examined two positively-selected regions, the gene ASPM and the human accelerated region (HAR) HAR1, for association with a set of speech and language related phenotypes under the hypothesis that these behaviors would be under positive selection. ASPM regulates brain size and is implicated in neuronal stem cell proliferation and differentiation. HAR1, a 118-bp region with an estimated 18 substitutions in the human lineage since human-chimpanzee divergence, bisects two RNA genes, HAR1F and HAR1R. It forms a stable structure and shows co-expression with reelin, a regulator of neocortex development. • ASPM • Two markers, rs6700180 and rs10737686, showed significant p-values across multiple measures (see table 2 and figure 1) • HAR1 • Four markers, rs6011605, rs6089838, rs7506966, and rs1884333, showed significant p-values across multiple language measures (see table 2 and figure 2) • Combined into a two SNP haplotype, rs6011605 and rs6089838, show significant association with spoken language (p=.00003) • Sequencing revealed two novel SNPs within the RNA genes HAR1F and HAR1R and next to HAR1 • chr20:61204086 (MAF 0.33) • chr20:61204162 (MAF 0.24) • Chr20:61204086, between rs6011605 and rs750696, only showed semi-significant association (p-value 0.08) for the Elision measure ASPM Table 2Significant SNP results with associating language measures HAR1 Figure 1 Gene structure, conservation, linkage disequilibrium, and significant single-point SNP results of language associations for the gene ASPM Methods Participants 581 second grade children who were members of a longitudinal cohort served as participants. The children were sampled from a larger population sample based on their spoken language status in kindergarten. Thirty-seven percent had poor spoken language (<10th percentile on two measures of language) in kindergarten, the remainder had normal language abilities. Phenotyping All children were administered a range of tasks that measured speech, language, reading and intelligence. With the exception of speech sound production ability which was obtained in kindergarten, the measures were obtained in 2nd grade. Table 1 describes these measures. Figure 2 Gene structure, conservation, linkage disequilibrium, and significant single-point SNP results of language associations for the region in and around HAR1 Conclusions • ASPM and HAR1, two regions showing human-specific evolution, associate with language measures • Additional studies looking at ASPM and HAR1 and their roles in neurodevelopment are necessary to define specific regions of involvement with cognition • ASPM • Two markers associate across multiple language measures • rs3762271 is a nonsynonymous SNP located in the 18th exon coding for leucine or isoleucine (C2648A) • rs10737686 is an intronic SNP located between exon 18 and 19, directly upstream of a LINE repeat (LIMA4) and near a highly conserved region • Both markers associate with separate measures, suggesting ASPM is involved in at least two cognitive pathways • HAR1 • This is first time RNA genes have been associated with language measures • Multiple markers associate with multiple cognitive measures, suggesting the loci surrounding HAR1 may be implicated in different cognitive pathways • This provides incentive to investigate other HARs (coding, intronic, and intergenic) and positively-selected regions for candidate loci involved in language Table 1. Behavioral phenotypes and specific tests used for phenotypes. Genotyping Saliva, blood, and/or cheek swabs were collected from nuclear families identified via a longitudinal language study. Samples were processed using standard protocols. Samples were typed using Taqman Pre-designed Genotyping Assays under standard reaction conditions. Five single nucleotide polymorphisms (SNPs) were genotyped for ASPM and seven around HAR1. Marker selection was based on haplotype block structure, minor allele frequency, and proximity to conserved regions. A case-control design based on language and reading composite scores was used to sequence 218 individuals at HAR1. Single SNP association was performed via linear regression, accounting for sibling and family effects by ASSOC in the S.A.G.E. software package. Haplotype analyses were performed using the Pedigree Disequilibrium Test (PDT) through Unphased. Acknowledgments Further Information keegan-kelsey@uiowa.edu http://genetics.uiowa.edu http://www.uiowa.edu/~clrc http://www.uiowa.edu This research was supported by grants DC00496 and DC02746 from the National Institute on Deafness and Other Communicative Disorders. We would also like to thank Katy Mueller, Marlea O’Brien, Rick Arenas, Kathy Frees, Juanita Limas, Marla Johnson, and Adela Mansilla for their work on this project.