RNAi Mechanism
340 likes | 593 Views
RNAi Mechanism. The Central Dogma. DNA (double-stranded). RNA (single-stranded). Protein. When good RNA goes bad. Viruses can make double- stranded RNA. When good RNA goes bad. Cells sense double- stranded RNA and activate a response called RNAi. Understanding how RNAi works is the

RNAi Mechanism
E N D
Presentation Transcript
The Central Dogma DNA (double-stranded) RNA (single-stranded) Protein
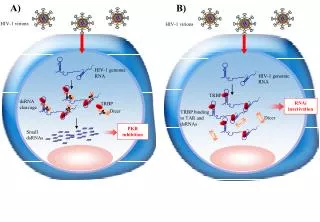
When good RNA goes bad.... Viruses can make double- stranded RNA
When good RNA goes bad.... Cells sense double- stranded RNA and activate a response called RNAi
Understanding how RNAi works is the key to using it as a genetic tool and for therapy
What is the mechanism of interference? RNAi in vitro... Cell free extract RNAi? Hannon Lab, Zamore Lab, Tuschl Lab, Sharp Lab
Dicer cuts dsRNA into short RNAs Vanhecke, D.; Janitz, M. Drug Discov. Today, 2005, 10, 205-225. Role for a bidentate ribonuclease in the initiation step of RNA interference Emily Bernstein, Amy A. Caudy, Scott M. Hammond & Gregory J. Hannon
dsRNA RNAi functions in many different organisms
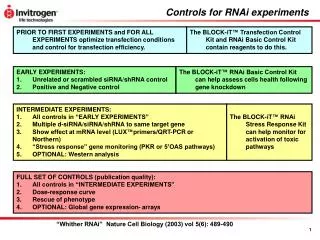
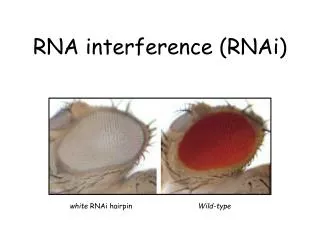
Dicer is required for RNAi wild-type GFP dsRNA GFP – germline transgene dcr-1-/-
siRNA is used as a template to direct cleavage of complementary mRNA by RISC RISC: RNA-inducing silencing complex
Argonaute proteins lie at the heart of RISC Ji-Joon Song Leemor Joshua-Tor
PIWI is an RNase H like domain Argonaute-PIWI RNase HI RNase HII Ji Joon Song Leemor Joshua-Tor
microRNAs regulate gene expression without cleavage
RNA or proteinlevel LIN-14, LIN-28 proteins LIN-41 protein LIN-29 protein let-7 RNA lin-4 RNA L1 stage L2 stage L3 stage L4 stage Adult stage Dicer stRNA precursor stRNA Translational repression small temporal RNAs (stRNAs) • lin-4 miRNA represses translation of lin-14 (L1 to L2 molt)
RNAi acts to regulate gene expression by two dicer-dependent mechanisms miRNA siRNA
Model for translational repression RISC AAAAAA......... M7GpppG RISC recognizes a target
Model for translational repression GW DCP RISC AAAAAA......... M7GpppG other Other proteins are also recruited – either along with RISC or later
Model for translational repression decapping to P-bodies GW DCP RISC AAAAAA......... M7GpppG other de-adenylation? (e.g. Giraldez, Belasco, Rivas) block translation? (e.g. Filipowicz) RISC may block through multiple mechanisms
Regulation of gene expression at the level of chromatin Sequence-independent linker histones: control DNA compaction and accessibility to trans-acting factors post-translational modifications of histone tails: control compaction of DNA and serve as docking sites for trans-acting factors Range: Can act at the level of a single gene, often acts over groups of genes and over larger domains (20-200kb), and can affect gene expression over an entire chromosome
Model for siRNA-dependent initiation of heterochromatic silencing by RITS
dsRNA-mediated silencing in various organisms: Multiple mechanisms that are Dicer-dependant Meister & Tuschl, 2004
dsRNA-mediated regulation may play a central role in the regulation of “gene networks” Levine and Davidson, PNAS, 2005 Ke, et al. Current Opinion in Chemical Biology 2003