Properties predicted by COSMO-RS-based QSAR
140 likes | 389 Views
Properties predicted by COSMO-RS-based QSAR. Dr. Karin Wichmann COSMO logic GmbH & Co. KG Burscheider Str. 515 D-51381 Leverkusen Germany Phone: +49-2171-731-683 Fax: +49-2171-731-689 Email: wichmann@cosmologic.de Web: http://www.cosmologic.de.

Properties predicted by COSMO-RS-based QSAR
E N D
Presentation Transcript
Properties predicted by COSMO-RS-based QSAR Dr. Karin Wichmann COSMOlogic GmbH & Co. KG Burscheider Str. 515 D-51381 Leverkusen Germany Phone: +49-2171-731-683 Fax: +49-2171-731-689 Email: wichmann@cosmologic.de Web: http://www.cosmologic.de
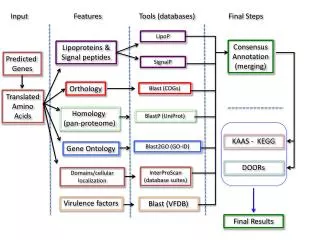
Property Prediction with the s-potential COSMOtherm allows for the prediction of the entire fluid phase equilibrium thermodynamics of almost arbitrary compounds, e.g: • Activity coefficients • Solubility • Partition coefficients between well characterized liquid phases The chemical potential and thermodynamic properties of liquids can easily be computed using the COSMO-RS s-potential mS(s): e.g. activity coefficients gXS:
Treatment of complex or chemically less well-defined matrices Some physiological and thermodynamic equilibria involve phases or matrices which are • Chemically not well defined or of unknown composition • Disordered • Not really liquid / amorphous E.g. phases like • Biological tissues, blood, brain, the intestinal resorption system • Soap, polymers, cotton • Solid adsorbents like activated carbon → s-potential cannot be calculated directly by COSMO-RS
Treatment of complex or chemically less well-defined matrices Chemically not well defined phases can be treated indirectly with the s-moments: The s-potential can be approximated very accurately by a set of functions MiX: s-moments MiX are defined with: And Hydrogen Bonding moments Mi,HBX M0X: molecular area M1X: negative total charge of compound X M2X: correlated with the screening energy for X M3X: “skewness” of the profile M-1/-2X or Mdon/accX: hydrogen bond capacity
Treatment of complex or chemically less well-defined matrices With the s-moments, • each phase / matrix can be characterized by a set of s-coefficients ciS. • Differences between two phases can be described by the same kind of expansion, with coefficients ciS,S‘ being the difference of the coefficients of the two phases. • Partition coefficients between two phases are generally expressed by a linear combination of the compounds s-moments in the form:
QSPR applications s-moments can be used as descriptors for linear QSAR/QSPR applications. Using COSMOtherms-moment descriptors it is possible to predict: • Partition behaviour between almost arbitrary phases, such as • CaCo2 cell permeability • logKBB blood-brain barrier partition • logKOC soil-water partitioning (environmental fate) • logKIA intestinal absorption • . . . and many more • Solubility of drugs in arbitrary solvents • Chemical potentials of crystal surfaces in solution → morphology of drugs
QSPR applications • Only five s-moments MiX are necessary to get a general QSPR description of solvents or biological matrices: • This is in correspondence with the general solvation theory of Abraham, which claims that solvent space is five-dimensional. • Intercorrelation of the COSMOtherms-moments and the Abraham set of solvent descriptors is possible* *Andreas M. Zissimos, Michael H. Abraham, Andreas Klamt, Frank Eckert, John Wood, J. Chem. Inf. Comput. Sci. 42 (2002) 1320
Blood-Brain Partitioning logBB = 0.0043 M0 - 0.017 M2 - 0.0031 M3 + 0.24 N = 103 rms = 0.41 R2 = 0.691 Exp. Data: Rose, K.; Hall, L. H.; Hall, L. M.; Kier, L. B. Modeling Blood-Brain Barrier Partitioning Using Topological Structure Descriptors, 2003. Elsevier MDL Web Site. http://www.mdl.com/solutions/white_papers/MDLQSARreprint.jsp (accessed Feb 13, 2004).
Human Serum Albumin Binding logKHSA = 0.0084 M0 - 0.017 M2 - 0.0014 M3 + 0.17 M3,acc + 0.24 N = 94 rms = 0.33 R2 = 0.671 Exp. Data: Kier, L. B.; Hall, L. H.; Hall, L. M. QSAR Modeling of Drug Binding to Protein. b-Lactam Serum Binding and Albumin Binding Affinity, 2002. Elsevier MDL Web Site. http://www.mdl.com/solutions/white_papers/ML233QM3_white_paper.jsp (accessed Oct 7, 2004).
Intestinal Absorption logKHSA = 0.0040 M0 - 0.0053 M2 - 0.0024 M3 - 0.11 M3,acc - 0.11 M3,don + 1.37
Soil Sorption logKOC = 0.016 M0 - 0.017 M2 - 0.046 M3 + 0.23 M3,acc – 0.33 M3,don + 0.41 N = 386 rms = 0.61 R2 = 0.72
Adsorption to Activated Carbon [Mehler, Peukert (TU München), Klamt; AICHE Journal. 48, 1093-1099 (2002)]
Model of Cytochrome P450 2D6 and Cytochrome P450 3A4 Dataset from Pharm. Res. 2003, 20, 1401. Log(IC50) = 2.96(0.56) – 0.0097(0.0018)*Area + 0.010(0.0067)*sig2 + 0.050(0.012)*sig3 - 0.42(0.089)*HBacc3 + 0.39(0.14)*HBdon3 Log(IC50) = 4.46(0.50) – 0.0093(0.0017)*Area - 0.0068(0.0078)*sig2 + 0.025(0.010)*sig3 + 0.033(0.128)*HBacc3 + 0.24(0.14)*HBdon3 Modelling inhibition of Cytochrome P450 3A4 using s-moment descriptors for 45 structurally diverse drugs. (2 outliers deleted.) Modelling inhibition of Cytochrome P450 2D6 using s-moment descriptors for a diverse set of 33 compounds from the NCI database.