Finding a maximum independent set in a sparse random graph
200 likes | 427 Views
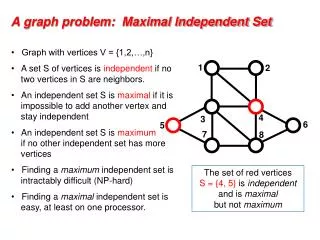
Finding a maximum independent set in a sparse random graph. Uriel Feige and Eran Ofek. Max Independent Set. Largest set of vertices that induce no edge. NP-hard, even to approximate. NP-hard on planar graphs.

Finding a maximum independent set in a sparse random graph
E N D
Presentation Transcript
Finding a maximum independent set in a sparse random graph Uriel Feige and Eran Ofek
Max Independent Set Largest set of vertices that induce no edge. • NP-hard, even to approximate. • NP-hard on planar graphs. • Polynomial time algorithms on “simple” graphs: trees (greedy), graphs of bounded treewidth (dynamic programming).
Complexity on most graphs • On random graphs, no polynomial time algorithm is known to find max-IS. • Holds for all densities except for extremely sparse or extremely dense graphs. • “Best” algorithmic lower bound – greedy. • “Best” upper bound – theta function.
Planted models • Model the case that a graph happens to have an exceptionally large IS. • Random graph with edge probability d/n. • All edges within a random set S of vertices are removed. • When d|S| > n log n, the set S is likely to be the maximum IS.
Some known results • When d=n/2, can find S of size [Alon, Krivelevich, Sudakov 1998]. • Can also certify maximality, and handle semirandom graphs [Feige, Krauthgamer 2000]. • When d=log n, can find S of size , even in semirandom graphs, up to the point when it becomes NP-hard [Feige, Kilian 2001]
Our results Allow d to be (a sufficiently large) constant. W.h.p., the random graph) has no independent set larger than n (log d)/d. Plant S of size We find max independent set in polynomial time. New aspect: S is not the max-IS. Complicates analysis.
S is not max-IS V-SS
Some related work • Many of the techniques in this area were initiated in work of Alon and Kahale (1997) on coloring. • Amin Coja-Oghlan (2005): finds a planted bisection in a sparse random graph. The min bisection is not the planted one. Amin’s algorithm is based on spectral techniques and certifies minimality.
Greedy algorithm • Select vertex i to put in solution (e.g., vertex i may be vertex of degree 0, degree 1, or of lowest degree). • Remove neighbors of i. • Repeat on G – i – N(i).
Simplify analysis 2-stage greedy • Select an independent set I. • Remove neighbors of I. • Finish off by exact algorithm. Last stage takes polynomial time if G-I-N(I) has “simple” structure.
Required properties of I Partition graph into Independent, Cover and Undecided. • No edge within I. • No edge between I and U. • Every vertex of C must have at least one neighbor in I. Note: U is then precisely V(G) – I – N(I).
How we select I Initialization. Threshold t = d(1 - |S|/2n) < d. • Put vertices of degree lower than t in I. • Put vertices of degree higher than t in C. Iteratively, move to U: • Vertices of I with neighbors in I or U. • Vertices of C with < 4 neighbors in I.
End of first step IC U
Theorems for planted model Lem: S highly correlated with max-IS. Lem: Low degree highly correlated with S. Thm: I is contained in max-IS. (Difficulty in proof: max-IS is not known not only to the algorithm, but also in analysis.) Thm: G(U) has simple structure.
Algorithm for G(U) Iteratively • Move vertices of degree 0 to I. • Move vertices of degree 1 to I, and their neighbors to C. Use exhaustive search to find maximum IS in each of the remaining connected components. Thm: CC of 2-core have size < O(log n).
Why did we consider 2-core? Asymmetry: vertices of S enter U more easily than vertices of V-S. A tree might have most its vertices from S. In a cycle, at least half the vertices must be from V-S. Easier to show that U has no large cycles then to show that has no large trees.
Conclusions • Planted model in sparse graphs, in which planted solution is not optimal. • Natural algorithm provably finds max-IS in planted model. (All difficulties are hidden in the analysis.) • Improve tradeoff between d and |S|. • Output matching upper bound on |max-IS|.