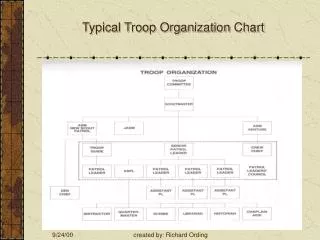
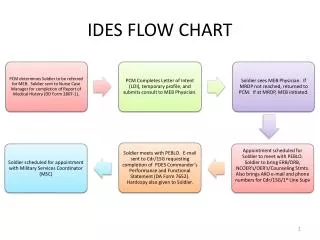
Flow Chart For a Typical MD Program
SMA5233 Particle Methods and Molecular Dynamics Lecture 6: Coarse-grained hybrid MD A/P Chen Yu Zong Tel: 6516-6877 Email: phacyz@nus.edu.sg http://bidd.nus.edu.sg Room 08-14, level 8, S16 National University of Singapore. Flow Chart For a Typical MD Program. Initialize Variables. Build

Flow Chart For a Typical MD Program
E N D
Presentation Transcript
SMA5233 Particle Methods and Molecular DynamicsLecture 6: Coarse-grained hybrid MDA/P Chen Yu ZongTel: 6516-6877Email: phacyz@nus.edu.sghttp://bidd.nus.edu.sgRoom 08-14, level 8, S16 National University of Singapore
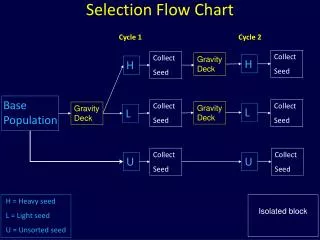
Flow Chart For a Typical MD Program Initialize Variables Build Model Define Material Set Boundary Conditions Start Initialize Variables • Define the positions of the atoms • Assign randomly generated velocities Integrate End
Flow Chart For a Typical MD Program Initialize Variables Build Model Define Material Set Boundary Conditions Start Build Model • Define the simulation domain • Build atoms to lattice Integrate End
Flow Chart For a Typical MD Program Initialize Variables Build Model Define Material Set Boundary Conditions Start Define Material • Specify interatomic potentials Integrate End
Flow Chart For a Typical MD Program Initialize Variables Build Model Define Material Set Boundary Conditions Start Set Boundary Conditions • Set initial temperature distribution • Specify thermodynamic controls Integrate End
Flow Chart For a Typical MD Program Initialize Variables Build Model Define Material Set Boundary Conditions Start Start • Compute the forces at time=0 • Set frequency of outputs Integrate End
Flow Chart For a Typical MD Program Initialize Variables Build Model Define Material Set Boundary Conditions Start Integrate • Compute the atomic trajectories • Compute other desired outputs Integrate End
Flow Chart For a Typical MD Program Initialize Variables Build Model Define Material Set Boundary Conditions Start End • Compute final thermodynamic outputs • Calculate program statistics Integrate End
Computation of material properties based on explicit treatment of atomic degrees of freedom Computationally expensive Too many degrees of freedom Only capable on small DNA duplexes Time duration in nanoseconds Fully Atomistic Simulations Limitations
Coarse-grained Model • DNA Sugar and Phosphate groups reduced to one molecule(bead) • Each DNA base is represented by one molecule(bead) Coarse-grained Model Fully Atomistic Model
Computationally less expensive Decreases degrees of freedom Advantages of the Coarse-grained Model • Allows for longer DNA duplexes • Time length up to microsecond Coarse-grained model DNA duplex Chemical structure of DNA duplex
Coarse-grained Model • One or multiple amino acids reduced to one molecule(bead)
Intra-Polymer Forces – Combinations Of the Following: • Lennard-Jones Repulsion • Stiff (Fraenkel) / Hookean Spring • Finitely-Extensible Non-linear Elastic (FENE) Spring
Intra-Polymer Forces – Combinations Of the Following: • Lennard-Jones Repulsion • Finitely-Extensible Non-linear Elastic (FENE) Spring
Intra-Polymer Forces (continued) • Marko-Siggia WormLike Chain Can be adjusted if M>2 (Underhill, Doyle 2004) Stiff: Schlijper, Hoogerbrugge, Manke, 1995 Hookean + Lennard-Jones: Nikunen, Karttunen, Vattulainen, 2003 FENE: Chen, Phan-Thien, Fan, Khoo, 2004