Microbiology Unknown Lab
Microbiology Unknown Lab. 0.1% dextrose 1.0% sucrose 1.0% lactose. TSI agar Triple Sugar Iron Agar. A B C D E. (a) Red/red (no sugar fermentation) (b) Control (c) Red/yellow (Glucose fermented but lactose and sucrose not fermented)

Microbiology Unknown Lab
E N D
Presentation Transcript
0.1% dextrose 1.0% sucrose 1.0% lactose TSI agarTriple Sugar Iron Agar A B C D E (a) Red/red (no sugar fermentation) (b) Control (c) Red/yellow (Glucose fermented but lactose and sucrose not fermented) (d) Yellow/yellow (Glucose fermented. Lactose and/or sucrose fermented) (e) Red/yellow with H2S
Penol Red broths A B C D E A (acid with gas)…..B (acid)……C (uninoculated control)……D (no recation)… …E (alkaline reaction)
O-F tests Control E. coli Pseudomonas Alcaligenes Fermenter Oxidizer Nonsaccharolytic
Decarboxylation uninoculated Positive Negative
Simmons Citrate A B C A: Positive…Enterobacter B: Negative…E. coli C: Control-uninoculated
Phenylalanine Deaminase Test neg pos
Urea Hydrolysis Positiveuninoculated Negative
Methyl Red Test A B A: Positive (E. coli) B: Negative (Enterobacter aerogenes)
Voges Proskauer Test A B A: Negative (E. coli) B: Positive (Enterobacter aerogenes)
Indole Test Negative Positive - +
Catalase Test + - Positive Negative
General Microbiology (BIOL 187) Laboratory Report (Unknown Identification) Page 1: Cover page Include your name, the unknown number and you final Genus and species identification Page 2: Test listing I want a (+) / (-) listing of each test in a vertical, easily scanned format Page 3: Discussion section Detail, in complete sentences, why your results justify your identification and rule out other possibilities. Page 4: About My Microbe section Research your microbe and give me a several paragraphs about your microbe. (Don't copy word-for-word from your textbook---This will be frowned upon!) Page 5: Dichotomous key Create a neat dichotomous key, which results in the correct separation of all 8 possible unknowns. This key should be legible and sized to occupy a complete sheet of paper (nothing microscopic is acceptable here) Page 6:Lab handout where you recorded your initial results Include the handout from lab where your recorded your data as you read the test reactions. Grading: 20 pts. Correct Identification 10 pts. Discussion section 7 pts. "About my microbe section" 8 pts. Dichotomous key 5 pts. Neatness, accuracy and completeness of report 50 points total
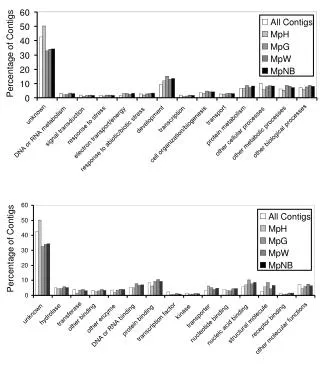
Differential Identification of Enterobacteriaceae by Biochemical Tests. How do I interpret the table above?? + means 90% or more of the strains are positive - means 90% or more of the strains are negative + or - means most strains are positive but a significant number are negative - or + means most strains are negative but a significant number are positive (+) means a positive result may be weak or delayed in its occurrence d means different results can be obtained. Tests marked with a d would be unreliable for identification purposes.
E.Coli Shigella flexneri Salmonella Citrobacter Proteus vulgaris Proteus mirabilis Enterobacter cloacae Enterobacter aerogenes - + Methyl Red Next…look for a test that distinguishes between the 2 methyl red negtive microbes
The lysine decarboxylase test distinguishes between these 2 microbes
E.Coli Shigella flexneri Salmonella Citrobacter Proteus vulgaris Proteus mirabilis Enterobacter cloacae Enterobacter aerogenes - + Methyl Red - + Lysine decarboxylase Enterobacter cloacae Enterobacter aerogenes NOW…look for a test that distinguishes between the 6 methyl red positive microbes.
The phenylalanine deaminase test has 4 negatives and 2 positives for the 6 microbes that are not yet “keyed-out” It would be a possibility for the next choice.
E.Coli Shigella flexneri Salmonella Citrobacter Proteus vulgaris Proteus mirabilis Enterobacter cloacae Enterobacter aerogenes - + Methyl Red - + - + Lysine decarboxylase Phenyalanine deaminase Enterobacter cloacae Enterobacter aerogenes E.Coli Shigella flexneri Salmonella Citrobacter (these 4 still need to be “keyed-out” from each other) Proteus vulgaris Proteus mirabilis (these 2 still need to be “keyed-out” from each other)