Donnie Berkholz
250 likes | 389 Views
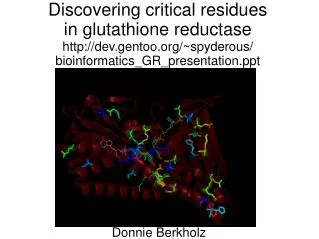
Discovering critical residues in glutathione reductase http://dev.gentoo.org/~spyderous/ bioinformatics_GR_presentation.ppt. Donnie Berkholz. What and How. Role Reduced thiols Oxidative stress DNA precursors H + transport Mechanism Flavoprotein NADPH Disulfide. Goals.

Donnie Berkholz
E N D
Presentation Transcript
Discovering critical residues in glutathione reductasehttp://dev.gentoo.org/~spyderous/bioinformatics_GR_presentation.ppt Donnie Berkholz
What and How • Role • Reduced thiols • Oxidative stress • DNA precursors • H+ transport • Mechanism • Flavoprotein • NADPH • Disulfide
Goals • Figure out the best programs and methods for this analysis • Search for unknown critical residues • Verify whether residues already thought critical are actually conserved • Check for potential differences in function and specificity among subfamilies (Podar et al.)
Multiple sequence alignments ClustalW Dialign-T Muscle ProbCons
ProbCons • “Probabilistic consistency” • Pair-HMM based • Three-way alignment consistency • Parameters derived from training • Maximized accuracy
How to find important residues? • Principal component analysis (PCA) • Each sequence becomes a vector • Successive dimensions grow less significant • Evolutionary trace and friends • Divide tree into groups, then check them • So, first we need trees
Trees • Maximum likelihood • ProML (PHYLIP) • Gamma distribution + invariant sites • Approximate with 5 rate categories • Bayesian • MrBayes • Gamma distribution + invariant sites • MCMC: Markov chain Monte Carlo • Mixed: sample with probability -> WAG • Try variable-rate models
ConSurf • Calculates evolutionary conservation (Bayesian) • Maps onto protein structure • Input flexibility • PDB -> seq. -> PSI-BLAST -> MSA -> NJ -> CS • Can't yet analyze subfamilies
What next? • Check for validity of tree model • Tree-determinant residues • Experimental functional determination
Summary • ProbCons is great for MSA's • Bayesian trees take forever, but they provide confidence values (no bootstrap!) • ConSurf maps sequence conservationonto protein structures • Supports catalytic hypothesis • New putative functional roles: • Interactions? F354+D22, D316+T321 • Binding: I26, R218 • Structure: H434 etc
References ClustalW: Chenna et al. NAR31: 3497 (2003). Muscle: Edgar. NAR32: 1792 (2004). Dialign-T: Morgenstern. NAR32: W33 (2004). ProbCons: Chuong et al. Genome Res.15: 330 (2005) Jalview: Clamp et al. Bioinform.12: 426 (2004). PHYLIP: Felsenstein. Distributed by author (2005). MrBayes: Ronquist and Huelsenbeck. Bioinform.19: 1572 (2003). ConSurf: Landau et al. NAR33: W299 (2005). PyMol: DeLano. www.pymol.org (2005).