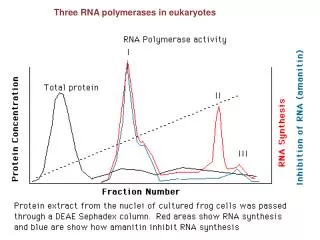
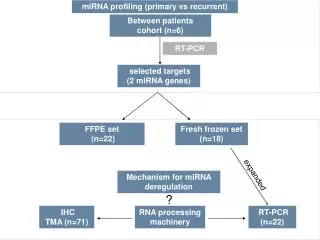
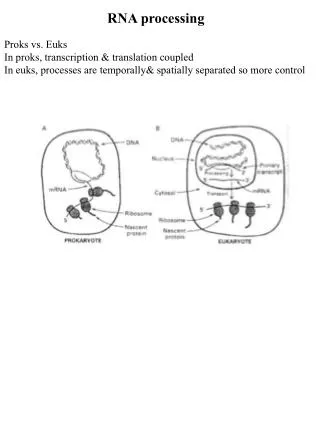
RNA processing in eukaryotes
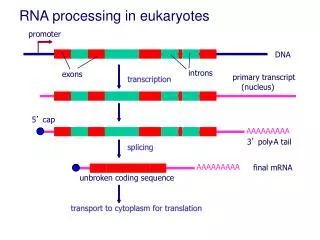
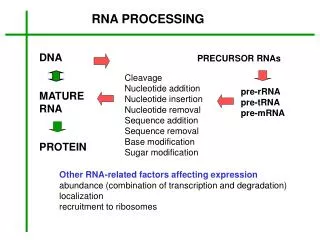
RNA processing in eukaryotes. promoter. DNA. introns. exons. primary transcript. transcription. transcription. (nucleus). 5 ’ cap. AAAAAAAAA. 3 ’ poly. -. A tail. splicing. splicing. AAAAAAAAA. final mRNA. unbroken coding sequence. transport to cytoplasm for translation.

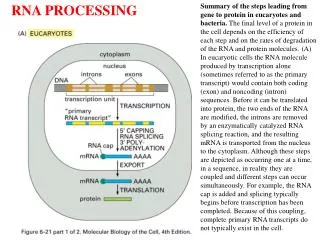
RNA processing in eukaryotes
E N D
Presentation Transcript
RNA processing in eukaryotes promoter DNA introns exons primary transcript transcription transcription (nucleus) 5’ cap AAAAAAAAA 3’ poly - A tail splicing splicing AAAAAAAAA final mRNA unbroken coding sequence transport to cytoplasm for translation transport to cytoplasm for translation
methylated guanine “backward” 5′ to 5′ linkage Not encoded in DNA Capping enzyme Recognition by ribosome 5′ cap 5′ AGACCUGACCAUACC
RNA processing in eukaryotes promoter DNA introns exons primary transcript transcription transcription (nucleus) 5’ cap AAAAAAAAA 3’ poly - A tail splicing splicing AAAAAAAAA final mRNA unbroken coding sequence transport to cytoplasm for translation transport to cytoplasm for translation
3′ poly(A) tail Poly(A) polymerase Add ~200 A’s Not in template Important for: Export of mRNA Initiation of Translation Stability of mRNA …UGGCAGACCUGACCA 3′ …UGGCAGACCUGACCAAAAAAAAAAAAAAAAAAAA
RNA processing in eukaryotes promoter DNA introns exons primary transcript transcription transcription (nucleus) 5’ cap AAAAAAAAA 3’ poly - A tail splicing splicing AAAAAAAAA final mRNA unbroken coding sequence transport to cytoplasm for translation transport to cytoplasm for translation
Splicing • Most genes interrupted by introns • Introns removed after transcription • Exons spliced together 5’ cap AAAAAAAAA 3’ poly - A tail splicing splicing AAAAAAAAA final mRNA unbroken coding sequence
Splicing snRNPs recognize exon-intron boundaries RNA + protein Cut and rejoin mRNA
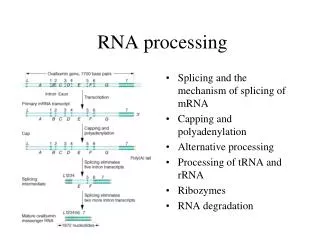
Splicing RPE65 mRNA in nucleus: 21,000 nt (14 exons) AAAAAAAAA splicing splicing AAAAAAAAA mature RPE65 mRNA in nucleus: 1,700 nt (8%)
Splicing Alternative splicing: >1 protein from one gene 27,000 human genes, but >100,000 proteins
Splicing Mutations affecting splicing can cause genetic disease: cystic fibrosis retinitis pigmentosa spinal muscular atrophy Prader-Willi syndrome Huntington disease spinocerebellar ataxia myotonic dystrophy Fragile-X syndrome Or produce genetic susceptibility to disease: lupus bipolar disorder schizophrenia myocardial infarction type I diabetes asthma cardiac hypertrophy multiple sclerosis autoimmune diseases elevated cholesterol
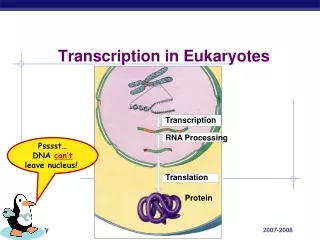
Gene expression summary Prokaryotes Eukaryotes DNA DNA transcription transcription mRNA pre-mRNA cytoplasm nucleus • capping • polyadenylation • splicing directly translated (even before being completely transcribed) protein mature mRNA • transport to cytoplasm • translation cytoplasm protein
Quick review of protein structure amino acids generic amino acid
Quick review of protein structure side chain gives chemical properties Non-polar (hydrophobic): Charged: Polar, not charged: Negative: Positive:
Quick review of protein structure polymer of amino acids = polypeptide ≈ protein methionine aspartate
Quick review of protein structure polymer of amino acids = polypeptide ≈ protein N- terminus C- terminus methionine aspartate peptide bond
polymer of amino acids = polypeptide ≈ protein Quick review of protein structure methionine aspartate glycine
polymer of amino acids = polypeptide ≈ protein Quick review of protein structure methionine aspartate glycine phenylalanine
polymer of amino acids = polypeptide ≈ protein Quick review of protein structure methionine aspartate glycine phenylalanine valine
polymer of amino acids = polypeptide ≈ protein Quick review of protein structure methionine aspartate glycine phenylalanine valine lysine
What holds folded proteins together? Hydrogen bonds Hydrophobic interactions Ionic bonds Disulfide bonds (covalent) …all determined by amino-acid sequence Quick review of protein structure
Primary (1°) structure Quick review of protein structure • Secondary (2°) structure alpha helix • Tertiary (3°) structure beta sheet • Quaternary (4°) structure hemoglobin L-isoaspartyl protein carboxyl methyltransferase
Translation Ribosome finds start codon within mRNA Genetic code determines amino acids Stop codon terminates translation start codon stop codon mRNA coding region 5′ 3′ translation protein NH3 COOH 5′ UTR 3′ UTR
Ribosome • Large ribonucleoprotein structure • E. coli: 3 rRNAs, 52 proteins • Two subunits: large and small large subunit RNA small subunit protein
Eukaryotic Translation • How does the ribosome find the correct start codon? • Small ribosome subunit binds 5′ cap • Scans to first AUG start codon stop codon mRNA AAAAAAAAA… 3′ coding region 5′ cap 5′ UTR 3′ UTR
Prokaryotic Translation • How does the ribosome find the correct start codon? • Small subunit binds Shine-Dalgarno sequence (RBS) • Positioned correctly for translation start codon stop codon mRNA coding region 5′ 3′ Shine-Dalgarno sequence or RBS (AGGAGG) 5′ UTR 3′ UTR
After finding start codon, use the genetic code: Shown as mRNA 5′ → 3′ the Genetic Code
Mechanics of Translation • Translation requires: • mature mRNA • ribosome • tRNAs • amino acids • accessory proteins
tRNA • Small RNAs (74-95 nt) made by transcription • Intramolecular base pairing • Anticodon complementary to mRNA codon anticodon
tRNA • “Charged” by specific aminoacyl tRNA synthetase
Initiation of Translation • Small ribosome subunit binds at start codon • Prokaryotes: Shine-Dalgarno sequence (RBS) • Eukaryotes: binds cap, scans mRNA 5′ AUG GAU GGG
Met Initiation of Translation • First tRNA (Met, anticodon CAU) joins complex 3' 5' UAC mRNA 5′ AUG GAU GGG
Met Initiation of Translation • Large ribosomal subunit joins 3' 5' UAC mRNA 5′ AUG GAU GGG
Met Initiation of Translation • P site holds tRNA with first aa • A site open for next tRNA 3' 5' UAC mRNA 5′ A P AUG GAU GGG
Met Asp Elongation • Next tRNA enters 3' 3' 5' 5' UAC CUA mRNA 5′ A P AUG GAU GGG
Met Asp Met Elongation • Peptidyl transferase forms peptide bond • Amino acid released from tRNA in P site mRNA 5′ 3' 5' 3' 5' UAC CUA AUG GAU GGG
Asp Met Elongation • Ribosome translocates one codon • First tRNA binds briefly in E site until translocation completes mRNA 5′ 3' 5' 3' 5' UAC CUA AUG GAU GGG
Asp Met Gly Elongation • Process repeats • Next tRNA can then enter the empty A site A P 3' 5' CCC mRNA 5′ 3' 5' CUA AUG GAU GGG
Lys Val Phe Val Asp Gly Gly Asp Met Ile Leu Leu Termination • Ribosome stops at stop codon • No matching tRNA • Release factor binds A Gln P RF 3' 5' GUC UUG CAG UAG
Translation complex dissociates Termination Lys Val Phe Val Met Gly Asp Asp Gly Ile Leu Leu Gln 3' 5' GUC UUG CAG UAG RF
Polyribosomes Next ribosome starts as soon as start codon is available N Ribosome AUG mRNA Stop C N Growing polypeptide RNA subunits released 3' 5' direction of ribosome movement 5' – 3' Released polypeptide
Operons • More than one gene on one mRNA • Prokaryotes only
Operons • More than one gene on one mRNA • Prokaryotes only
Protein Synthesis Pathways • Free ribosomes • Ribosomes bound to RER