Chapter 21 RNA Splicing and Processing
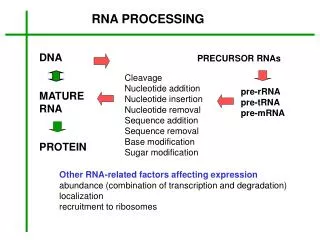
Chapter 21 RNA Splicing and Processing. 21.1 Introduction. pre-mRNA – The nuclear transcript that is processed by modification and splicing to give an mRNA. RNA splicing – The process of excising introns from RNA and connecting the exons into a continuous mRNA. 21.1 Introduction.

Chapter 21 RNA Splicing and Processing
E N D
Presentation Transcript
21.1 Introduction • pre-mRNA –The nuclear transcript that is processed by modification and splicing to give an mRNA. • RNA splicing – The process of excising introns from RNA and connecting the exons into a continuous mRNA.
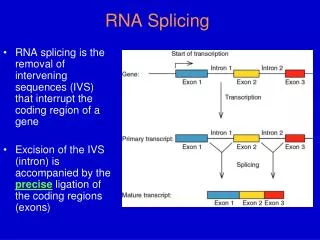
21.1 Introduction Figure 21.01: RNA is modified in the nucleus by additions to the 5’ and 3’ ends and by splicing to remove the introns.
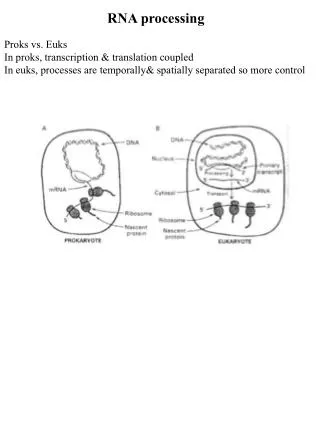
21.1 Introduction • heterogeneous nuclear RNA (hnRNA) – RNA that comprises transcripts of nuclear genes made by RNA polymerase II; it has a wide size distribution and low stability. • hnRNP – The ribonucleoprotein form of hnRNA (heterogeneous nuclear RNA), in which the hnRNA is complexed with proteins. • Pre-mRNAs are not exported until processing is complete; thus they are found only in the nucleus.
21.2 The 5′ End of Eukaryotic mRNA Is Capped • A 5′ cap is formed by adding a G to the terminal base of the transcript via a 5′–5′ link. • The capping process takes place during transcription and may be important for release from pausing of transcription. Figure 21.02: The cap blocks the 5’ end of mRNA and is methylated at several positions.
21.2 The 5′ End of Eukaryotic mRNA Is Capped • The 5′ cap of most mRNA is monomethylated, but some small noncoding RNAs are trimethylated. • The cap structure is recognized by protein factors to influence mRNA stability, splicing, export, and translation.
21.3 Nuclear Splice Sites Are Short Sequences • Splice sites are the sequences immediately surrounding the exon–intron boundaries. They are named for their positions relative to the intron. • The 5′ splice site at the 5′ (left) end of the intron includes the consensus sequence GU. • The 3′ splice site at the 3′ (right) end of the intron includes the consensus sequence AG.
21.3 Nuclear Splice Sites Are Short Sequences • The GU-AG rule (originally called the GT-AG rule in terms of DNA sequence) describes the requirement for these constant dinucleotides at the first two and last two positions of introns in pre-mRNAs. Figure 21.03: The ends of nuclear introns are defined by the GU-AG rule.
21.3 Nuclear Splice Sites Are Short Sequences • There exist minor introns relative to the major introns that follow the GU-AG rule. • Minor introns follow a general AU-AC rule with a different set of consensus sequences at the exon–intron boundaries.
21.4 Splice Sites Are Read in Pairs • Splicing depends only on recognition of pairs of splice sites. • All 5′ splice sites are functionally equivalent, and all 3′ splice sites are functionally equivalent. • Additional conserved sequences at both 5′ and 3′ splice sites define functional splice sites among numerous other potential sites in the pre-mRNA.
21.4 Splice Sites Are Read in Pairs Figure 21.04: Splicing junctions are recognized only in the correct pairwise combinations.
21.5 Pre-mRNA Splicing Proceeds Through a Lariat • Splicing requires the 5′ and 3′ splice sites and a branch site just upstream of the 3′ splice site. • The branch sequence is conserved in yeast but less well conserved in multicellular eukaryotes. • A lariat is formed when the intron is cleaved at the 5′ splice site, and the 5′ end is joined to a 2′ position at an A at the branch site in the intron.
21.5 Pre-mRNA Splicing Proceeds Through a Lariat • The intron is released as a lariat when it is cleaved at the 3′ splice site, and the left and right exons are then ligated together. Figure 21.05: Splicing occurs in two stages. First the 5’ exon is cleaved off, and then it is joined to the 3’ exon.
21.6 snRNAs Are Required for Splicing • small cytoplasmic RNAs (scRNA; scyrps) – RNAs that are present in the cytoplasm (and sometimes are also found in the nucleus). • small nuclear RNA (snRNA; snurps) – One of many small RNA species confined to the nucleus; several of them are involved in splicing or other RNA processing reactions. • small nucleolar RNA (snoRNA) – A small nuclear RNA that is localized in the nucleolus.
21.6 snRNAs Are Required for Splicing • The five snRNPs involved in splicing are U1, U2, U5, U4, and U6. • Together with some additional proteins, the snRNPs form the spliceosome. Figure 21.07: The spliceosome is ~12 MDa. Five snRNPs account for almost half of the mass.
21.6 snRNAs Are Required for Splicing • All the snRNPs except U6 contain a conserved sequence that binds the Sm proteins that are recognized by antibodies (anti-SM) generated in autoimmune disease. • splicing factor – A protein component of the spliceosome that is not part of one of the snRNPs. • transesterification – A reaction that breaks and makes chemical bonds in a coordinated transfer so that no energy is required.
21.7 Commitment of Pre-mRNA to the Splicing Pathway • U1 snRNP initiates splicing by binding to the 5′ splice site by means of an RNA–RNA pairing reaction. • The commitment complex (or E complex) contains U1 snRNP bound at the 5′ splice site and the protein U2AF bound to a pyrimidine tract between the branch site and the 3′ splice site.
21.7 Commitment of Pre-mRNA to the Splicing Pathway Figure 21.10: The commitment (E) complex formation.
21.7 Commitment of Pre-mRNA to the Splicing Pathway • In cells of multicellular eukaryotes, SR proteins play an essential role in initiating the formation of the commitment complex. • Pairing splice sites can be accomplished by intron definition or exon definition.
21.7 Commitment of Pre-mRNA to the Splicing Pathway Figure 21.11: There are two routes for initial recognition of 5’ and 3’ splice sites by either intron definition or exon definition.
21.8 The Spliceosome Assembly Pathway • The commitment complex progresses to pre-spliceosome (the A complex) in the presence of ATP. • Recruitment of U5 and U4/U6 snRNPs converts the A complex to the mature spliceosome (the B1 complex). • The B1 complex is next converted to the B2 complex in which U1 snRNP is released to allow U6 snRNA to interact with the 5′ splice site.
21.8 The Spliceosome Assembly Pathway • When U4 dissociates from U6 snRNP, U6 snRNA can pair with U2 snRNA to form the catalytic active site. • Both transesterification reactions take place in the activated spliceosome (the C complex). • The splicing reaction is reversible at all steps. Figure 21.12: The splicing reaction proceeds through discrete stages.
21.9 An Alternative Spliceosome Uses Different snRNPs to Process the Minor Class of Introns • An alternative splicing pathway uses another set of snRNPs that comprise the U12 spliceosome. • The target introns are defined by longer consensus sequences at the splice junctions rather than strictly according to the GU-AG or AU-AC rules. • Major and minor spliceosomes share critical protein factors, including SR proteins.
21.10 Pre-mRNA Splicing Likely Shares the Mechanism with Group II Autocatalytic Introns • Group II introns excise themselves from RNA by an autocatalytic splicing event (autosplicing or self-splicing). • The splice junctions and mechanism of splicing of group II introns are similar to splicing of nuclear introns. • A group II intron folds into a secondary structure that generates a catalytic site resembling the structure of U6-U2-nuclear intron.
21.10 Pre-mRNA Splicing Likely Shares the Mechanism with Group II Autocatalytic Introns Figure 21.15: Three classes of splicing reactions proceed by two transesterifications.
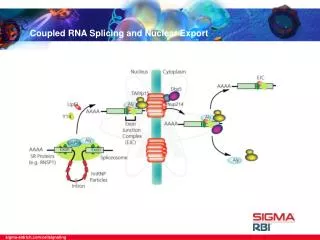
21.11 Splicing Is Temporally and Functionally Coupled with Multiple Steps in Gene Expression • Splicing can occur during or after transcription. • The transcription and splicing machineries are physically and functionally integrated. • Splicing is connected to mRNA export and stability control. • exon junction complex (EJC) – A protein complex that assembles at exon–exon junctions during splicing and assists in RNA transport, localization, and degradation.
21.11 Splicing Is Temporally and Functionally Coupled with Multiple Steps in Gene Expression Figure 21.18: The EJC (exon junction complex) is deposited near the splice junction as a consequence of the splicing reaction.
21.11 Splicing Is Temporally and Functionally Coupled with Multiple Steps in Gene Expression • Splicing in the nucleus can influence mRNA translation in the cytoplasm. • nonsense-mediated mRNA decay (NMD) – A pathway that degrades an mRNA that has a nonsense mutation prior to the last exon. Figure 21.20: The EJC complex couples splicing with NMD.
21.12 Alternative Splicing Is a Rule, Rather Than an Exception, in Multicellular Eukaryotes • Specific exons or exonic sequences may be excluded or included in the mRNA products by using alternative splicing sites. • Alternative splicing contributes to structural and functional diversity of gene products.
21.12 Alternative Splicing Is a Rule, Rather Than an Exception, in Multicellular Eukaryotes Figure 21.21: Different modes of alternative splicing.
21.12 Alternative Splicing Is a Rule, Rather Than an Exception, in Multicellular Eukaryotes • Sex determination in Drosophila involves a series of alternative splicing events in genes encoding successive products of a pathway.
21.12 Alternative Splicing Is a Rule, Rather Than an Exception, in Multicellular Eukaryotes Figure 21.23: Sex determination in D. melanogaster involves a pathway in which different splicing events occur in females.
21.13 Splicing Can Be Regulated by Exonic and Intronic Splicing Enhancers and Silencers • Alternative splicing is often associated with weak splice sites. • Sequences surrounding alternative exons are often more evolutionarily conserved than sequences flanking constitutive exons. • Specific exonic and intronic sequences can enhance or suppress splice site selection.
21.13 Splicing Can Be Regulated by Exonic and Intronic Splicing Enhancers and Silencers Figure 21.24: Exonic and intronic sequences can modulate the splice site selection by functioning as splicing enhancers or silencers.
21.13 Splicing Can Be Regulated by Exonic and Intronic Splicing Enhancers and Silencers • The effect of splicing enhancers and silencers is mediated by sequence-specific RNA binding proteins, many of which may be developmentally regulated and/or expressed in a tissue-specific manner. • The rate of transcription can directly affect the outcome of alternative splicing. Figure 21.25: The Nova and Fox families of RNA binding proteins can promote or suppress splice site selection in a context dependent fashion.
21.14 trans-Splicing Reactions Use Small RNAs • Splicing reactions usually occur only in cis between splice sites on the same molecule of RNA. • trans-splicing occurs in trypanosomes and worms where a short sequence (SL RNA) is spliced to the 5′ ends of many precursor mRNAs. • SL RNAs have a structure resembling the Sm-binding site of U snRNAs. Figure 21.26: Splicing usually occurs only in cis between exons carried on the same physical RNA molecule.
21.15 The 3′ Ends of mRNAs Are Generated by Cleavage and Polyadenylation • The sequence AAUAAA is a signal for cleavage to generate a 3′ end of mRNA that is polyadenylated. • The reaction requires a protein complex that contains a specificity factor, an endonuclease, and poly(A) polymerase. • The specificity factor and endonuclease cleave RNA downstream of AAUAAA. Figure 21.29: The sequence AAUAAA is necessary for cleavage to generate a 3’ end for polyadenylation.
21.15 The 3′ Ends of mRNAs Are Generated by Cleavage and Polyadenylation • The specificity factor and poly(A) polymerase add ~200 A residues processively to the 3′ end. • The poly(A) tail controls mRNA stability and influences translation. • Cytoplasmic polyadenylation plays a role in Xenopus embryonic development. Figure 21.30: The 3’ processing complex consists of several activities.
21.16 The 3′ mRNA End Processing Is Critical for Termination of Transcription • There are various ways to end transcription by different RNA polymerases. • The mRNA 3′ end formation signals termination of Pol II transcription. Figure 21.31: Transcription by Pol I and Pol III uses specific terminators to end transcription.
21.17 The 3′ End Formation of Histone mRNA Requires U7 snRNA • The expression of histone mRNAs is replication dependent and is regulated during the cell cycle. • Histone mRNAs are not polyadenylated; their 3′ ends are generated by a cleavage reaction that depends on the structure of the mRNA.
21.17 The 3′ End Formation of Histone mRNA Requires U7 snRNA • The cleavage reaction requires the SLBP to bind to a stem-loop structure and the U7 snRNA to pair with an adjacent single-stranded region. • The cleavage reaction is catalyzed by a factor shared with the polyadenylation complex.
21.17 The 3′ End Formation of Histone mRNA Requires U7 snRNA Figure 21.33: Generation of the 3’ end of histone H3 mRNA depends on a conserved hairpin and a sequence that base pairs with U7 snRNA.
21.18 tRNA Splicing Involves Cutting and Rejoining in Separate Reactions • RNA polymerase III terminates transcription in a poly(U)4 sequence embedded in a GC-rich sequence. • tRNA splicing occurs by successive cleavage and ligation reactions. Figure 21.34: The intron in yeast tRNAPhe base pairs with the anticodon to change the structure of the anticodon arm.
21.18 tRNA Splicing Involves Cutting and Rejoining in Separate Reactions • An endonuclease cleaves the tRNA precursors at both ends of the intron. • Release of the intron generates two half-tRNAs with unusual ends that contain 5′ hydroxyl and 2′–3′ cyclic phosphate. Figure 21.36: The 3’ and 5’ cleavages in S. cerevisiae pre-tRNA are catalyzed by different subunits of the endonuclease.
21.18 tRNA Splicing Involves Cutting and Rejoining in Separate Reactions • The 5′–OH end is phosphorylated by a polynucleotide kinase, the cyclic phosphate group is opened by phosphodiesterase to generate a 2′–phosphate terminus and 3′–OH group, exon ends are joined by an RNA ligase, and the 2′–phosphate is removed by a phosphatase.
21.18 tRNA Splicing Involves Cutting and Rejoining in Separate Reactions Figure 21.38: Splicing of tRNA requires separate nuclease and ligase activities.
21.19 The Unfolded Protein Response Is Related to tRNA Splicing • Ire1 is an inner nuclear membrane protein with its N-terminal domain in the ER lumen and its C-terminal domain in the nucleus; the C-terminal domain exhibits both kinase and endonuclease activities. • Binding of an unfolded protein to the N-terminal domain activates the C-terminal endonuclease by autophosphorylation.
21.19 The Unfolded Protein Response Is Related to tRNA Splicing Figure 21.39: The unfolded protein response occurs by activating special splicing of HAC1 mRNA to produce a transcription factor recognizing the UPRE.
21.19 The Unfolded Protein Response Is Related to tRNA Splicing • The activated endonuclease cleaves HAC1 (Xbp1 in vertebrates) mRNA to release an intron and generate exons that are ligated by a tRNA ligase. • Only spliced HAC1 mRNA can be translated to a transcription factor that activates genes encoding chaperones that help to fold unfolded proteins. • Activated Ire1 induces apoptosis when the cell is overstressed by unfolded proteins.
21.20 Production of rRNA Requires Cleavage Events and Involves Small RNAs • RNA polymerase I terminates transcription at an 18-base terminator sequence. • The large and small rRNAs are released by cleavage from a common precursor rRNA; the 5S rRNA is separately transcribed. Figure 21.40: Mature eukaryotic rRNAs are generated by cleavage and trimming events from a primary transcript.