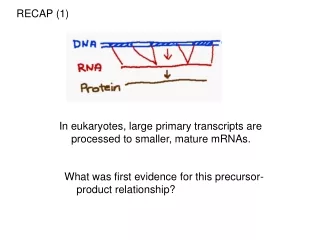
Eukaryotic mRNA Transcripts are Processed
Cytoplasm. Nuclear pores. AAAAAA. AAAAAA. DNA. Transcription. RNA. RNA Processing. G. G. mRNA. Export. Nucleus. Eukaryotic mRNA Transcripts are Processed. Transcription Start Site. 3’ UTR. 5’ UTR. Introns. 5’. 3’. Int. 1. Int. 2. Exon 1. Exon 2. Exon 3. Promoter/

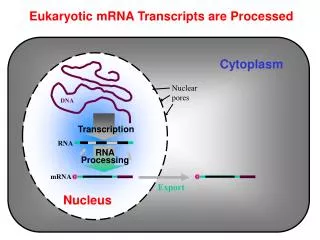
Eukaryotic mRNA Transcripts are Processed
E N D
Presentation Transcript
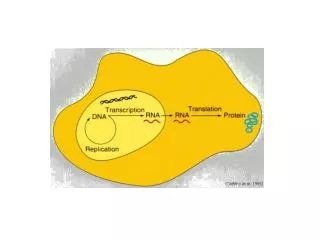
Cytoplasm Nuclear pores AAAAAA AAAAAA DNA Transcription RNA RNA Processing G G mRNA Export Nucleus Eukaryotic mRNA Transcripts are Processed
Transcription Start Site 3’ UTR 5’ UTR Introns 5’ 3’ Int. 1 Int. 2 Exon 1 Exon 2 Exon 3 Promoter/ Control Region Terminator Sequence Exons 5’ 3’ Int. 1 Int. 2 Exon 1 Exon 2 Exon 3 Unprocessed RNA Transcript A “Simple” Eukaryotic Gene Transcription
5’ UTR 3’ UTR AAAAA Protein Coding Region G Exon 1 Exon 2 Exon 3 5’ Cap 3’ Poly A Tail Int. 1 Int. 2 Processing of eukaryotic mRNA • RNA processing achieves three things: 1) Removal of Introns 2) Addition of a 5’ cap 3) Addition of a 3’ tail • The mRNA then moves out of the nucleus and is translated in the cytoplasm.
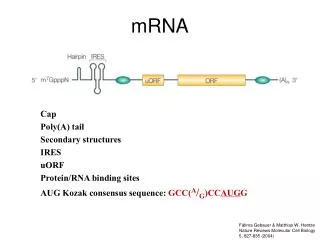
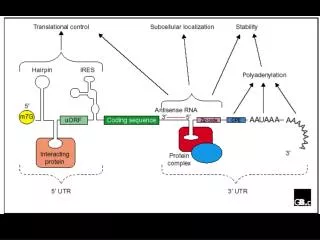
Translational control Structure of eukaryotic and prokaryotic mRNAs:
Model of eukaryotic ribosome • rRNAs are believed to play a catalytic role in protein synthesis. • After removal of 95% of the ribosomal proteins, the 60S subunit can catalyze formation of peptide bonds. • Ribosomal proteins are now believed to help fold the rRNAs properly and to position the tRNAs.
Small & large ribosomal subunits. • A Binding site for the mRNA is present on small subunit. • Two binding sites (P & A) bind tRNAs on large subunit. • P site – holds the tRNA carrying the growing polypeptide chain. • A site – holds the tRNA with the next AA to be added. • Ribosomes hold the mRNA and tRNAs together and connect the amino acids at the A site to the growing polypeptide. Ribosome Structure
Structure of tRNA • Aligns each amino acid with the corresponding codon • • 70-80 nt long • • 3’ end has the 5’- CCA sequence to which aa are linked • • The opposite end contains the anticodon loop • •Contains modified bases
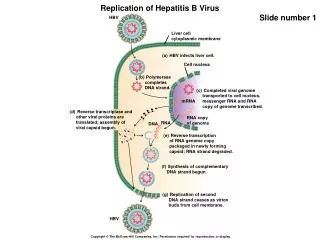
Many RNA Viruses have capped genomic RNAs similar to eukaryotic host mRNAs • Most eukaryotic mRNAs are capped at the 5’ end during nuclear processing. • The terminal 5’ phosphate is first removed by a 5’ triphosphatase. • Guanyltransferase transfers GMP from GTP to the 5’ end of the mRNA to add the GpppN cap structure. • The 5’ terminal inverted G residue is then modified by methylation. • Many RNA viruses replicate in the cytoplasm and must use a viral dependent capping mechanism supplied by the RNA-Dependent-RNA Polymerase. • The Cap structure, m7GpppN, is most common in viral and mammalian mRNAs.
Three distinct stages of translation • Initiation • •Rate limiting step • •Requires hydrolysis of ATP and GTP • •Results in formation of a complex containing the mRNA, the ribosome and the initiator Met-tRNA • A.5’ end (Cap) dependent initiation • • The initiation complex binds to the 5’ cap structure and scans in a 5’ to 3’ direction until initiating AUG is encountered • B. Internal ribosome entry • Initiation complex binds upstream of initiation codon
5’ end (cap) dependent initiation: • • The first step is the recognition of the 5’ cap by eIF4F, which consists of three proteins, eIF4E, eIF4G and eIF4A. • Cap binding protein, eIF4E, binds to cap • • The N-terminus of eIF4G binds eIF4E and the C-terminus binds eIF4A • • The 40S subunit binds to eIF4G via eIF3
Cap-Dependent Initiation of Protein Synthesis in Eukaryotes An initiation complex forms at the cap with the 40S ribosomal subunit and other translation initiation factors. The 40S complex then scans down the 5’ untranslated region to the first AUG codon. A GTP hydrolysis step by eIF5 triggers GDP binding of eIF2 and release of initiation proteins. The 60S subunit joins the complex and the 80S ribosome initiates translate the ORF.
Elongation eEF1a GTP Ribosome selects aminoacylated tRNA eEF1a and GTP are bound to aminoacylated tRNA Ribosome catalyzes formation of a peptide bond Translocation is dependent on eEF2 and GTP hydrolysis Many ribosomes may translate mRNAs simultaneously on the same strand. P A eEF2 GTP
Termination • Translation is terminated • at one of three stop codons (UAA, UAG & UGA). • • Termination codon at the A site is recognized by the release factor instead of a tRNA • • The release factor binds the termination codon • • The peptide chain is then released followed by dissociation of the tRNA and the ribosome
Closed loop model: • • The 5’ end dependent initiation is stimulated by the poly(A) binding protein Pabp1p, which interacts with eIF4G • • This interaction circularizes the mRNA and facilitates formation of the initiation complex • Mechanism to ensure that only intact mRNA is translated
5’end (cap) independent initiation: Poliovirus • • mRNAs of picornaviruses lack 5’ cap • Ribosomes bind internally rather than at the mRNA 5’ end • 5’ end of poliovirus RNA promotes internal binding of 40S subunit at internal ribosome entry site (IRES) • In poliovirus infected cells eIF4G is cleaved, inactivating translation of cellular mRNAs • The initiation in the IRES does not depend on the presence of a cap structure, but requires C-terminal fragment of eIF4G to recruit the 40S subunit through interaction with eIF3.
Five different types of IRES sequences are found on viral RNAs: Type I-entero and rhinoviruses (poliovirus) Initiation codon is located past the 3’ end of the IRES 40S binds to IRES scans to AUG Type II-cardio and apthoviruses (EMCV) Initiation codon is at the 3’ end of the IRES 40S binds at or near AUG no scanning occurs Type III- hepatitis A virus initiation codon is located past the 3’ end of the IRES requires all of initiation proteins, including eIF4E Type IV- hepatitis C virus The 3’ end of the hepatitis C virus IRES extends beyond the AUG codon Type V-cricket paralysis virus IRES ends at the initiation codon, although it is not an AUG codon, no initiation factors are required initiation codon is placed at the A site instead of the P site
Four types of IRES: Hepatitis C polivirus Encephelomyocarditis virus (EMCV) Cricket paralysis virus
Hepatitis C virus IRES: • 40S ribosomal subunit binds directly to Hepatitis C virus IRES in the absence of most initiation factors • A dramatic change in the conformation of the 40S subunit occurs when it binds Hepatitis C virus IRES setting the AUG at the P site • eIF3 also binds to the Hepatitis C virus IRES
Viral translation strategies Polyprotein synthesis Picornaviruses- Entire (+) sense RNA genome is translated into a single large polyprotein. Processing is carried out by two virus encoded proteases 2A pro and 3C pro. Flaviviruses- Viral precursor proteins are processed by cellular proteases. The (+) sense RNA genome is translated into a polyprotein precursor processed by viral serine protease and by host signal peptidase. Potyvirus group of plant viruses- Potato virus Y and tobacco etch virus contain a (+) sense genome RNA of around 10,000 bases which has a single open reading frame. This polyprotein is processed by viral encoded proteases.
Polyprotein processing in enteroviruses and flaviviruses Poliovirus: Flavivirus:
Leaky scanning Although majority of eukaryotic mRNAs are monocistronic, some viral mRNAs encode overlapping reading frames. The first start site is in a poor context, some ribosomes can bypass it and initiate at the second AUG, which has a better context. This will result in translation of two different proteins.
Reinitiation Rare in eukaryotes, but very common in prokaryotic cellular and viral mRNAs. Some eukaryotic mRNAs contain upstream AUG codons that terminate before the downstream reading frame. The upstream open reading frames may be translated, with reinitiation occurring at the downstream open reading frame.
Reinitiation of translation in influenza B virus In influenza B virus mRNA, M2 initiation codon is part of the termination codon for M1 protein. M2 synthesis is not efficient and dependent on M1 synthesis
Suppression of termination Suppression of termination occurs during translation of may viral mRNAs as a means of generating a second protein with extended carboxy terminus. In retroviruses, gag and pol genes are encoded by a single mRNA and separated by an amber termination codon UAG. Translational suppression of the amber codon allows synthesis of the gag pol precursor. A similar strategy is used by tobacco mosaic virus to translate its replicase proteins. Translation suppression is mediated by suppressor tRNAs that can recognize termination codons and insert a specific amino acid. The nucleotide sequence 3’ of the termination codon also plays an important role in the efficiency of translational suppression.
Suppression of termination codons in alphaviruses and retroviruses Pseudonot alphavirus retrovirus
Ribosomal frameshifting Ribosomal frameshifting is a process in which ribosomes move to a different reading frame and continue translation in that reading frame. It was discovered in cells infected with Rous sarcoma virus and has since been described for many other viruses including other retroviruses, (+) strand RNA viruses and herpes simplex virus. Requires a “slippery” sequence X-XXY-YYZ (in Rous sarcoma virus A-AAU-UUA) and an RNA secondary structure called a pseudoknot five to eight nucleotides downstream. Two tRNAs in the zero reading frame slip back by one nucleotide to the –1 phase and each tRNA base pairs with the mRNA in the first two nucleotides of each codon. As a result of the frameshift a Gag-pol fusion is produced at about 5% of the level of Gag protein.
Ribosomal frameshifting Rous sarcoma virus mRNA encodes Gag and Pol proteins that overlap in a –1 reading frame
REGULATION OF TRANSLATION DURING VIRAL INFECTION Interferons are produced in response to viral infection as part of the rapid innate immune response Interferons bind to cell surface receptors and activate transcription of antiviral genes Two interferon induced genes encode RNase L and protein kinase RNA-activated (Pkr) RNase L degrades RNA Pkr phosphorylates eIF2a, inhibiting translation initiation Pkr is a serine threonine kinase composed of an N-terminal regulatory domain and a C-terminal catalytic domain Pkr is activated by the binding of dsRNA to two dsRNA binding motifs at the N-terminus of the protein.
Viral regulation of Pkr Viruses use at least five different mechanisms to block Pkr activation or to stop activated Pkr from inhibiting translation • inhibition of dsRNA binding- adenovirus VA RNA binds Pkr blocks its activation by dsRNA • vaccinia virus E3L protein sequesters ds RNA • inhibition of Pkr dimerization influenza virus P58 hepatitis C virus NS5A • inhibitors of kinase function- vaccinia virus K3L protein has homology to N- terminus of eIF2-a