Intracellular Networks
C. E. N. T. E. R. F. O. R. I. N. T. E. G. R. A. T. I. V. E. B. I. O. I. N. F. O. R. M. A. T. I. C. S. V. U. Intracellular Networks. (2) Intracellular Network Behaviour Protein Function Prediction. Networks.

Intracellular Networks
E N D
Presentation Transcript
C E N T E R F O R I N T E G R A T I V E B I O I N F O R M A T I C S V U Intracellular Networks (2) Intracellular Network Behaviour Protein Function Prediction
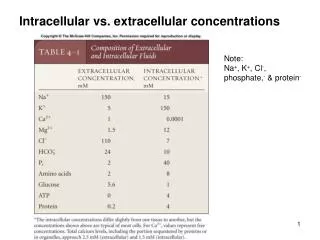
Networks "The thousands of components of a living cell are dynamically interconnected, so that the cell’s functional properties are ultimately encoded into a complex intracellular web [network] of molecular interactions." "This is perhaps most evident with cellular metabolism, a fully connected biochemical network in which hundreds of metabolic substrates are densely integrated through biochemical reactions." (Ravasz E, et al.)
TF Ribosomal proteins
Network Evolution Human Yeast This pathway diagram shows a comparison of pathways in (left) Homo sapiens(human) and (right)Saccharomycescerevisiae(baker’s yeast). Changes in controlling enzymes (square boxes in red) and the pathway itself have occurred (yeast has one altered (‘overtaking’) path in the graph)
The citric-acid cycle http://en.wikipedia.org/wiki/Krebs_cycle
The citric-acid cycle Fig. 1. (a) A graphical representation of the reactions of the citric-acid cycle (CAC), includingthe connections with pyruvate and phosphoenolpyruvate, and the glyoxylate shunt.When there are two enzymes that are not homologous to each other but that catalyse thesame reaction (non-homologous gene displacement), one is marked with a solid line andthe other with a dashed line. The oxidative direction is clockwise. The enzymes with theirEC numbers are as follows: 1,citrate synthase (4.1.3.7); 2, aconitase (4.2.1.3); 3, isocitratedehydrogenase (1.1.1.42); 4, 2-ketoglutarate dehydrogenase(solid line; 1.2.4.2 and2.3.1.61) and 2-ketoglutarate ferredoxin oxidoreductase (dashed line; 1.2.7.3); 5, succinyl-CoA synthetase (solid line; 6.2.1.5) or succinyl-CoA–acetoacetate-CoA transferase(dashed line; 2.8.3.5); 6, succinate dehydrogenase or fumarate reductase (1.3.99.1); 7,fumarase (4.2.1.2) class I (dashed line) and class II (solid line); 8, bacterial-type malatedehydrogenase (solid line) or archaeal-type malate dehydrogenase (dashed line)(1.1.1.37); 9, isocitrate lyase (4.1.3.1); 10, malate synthase (4.1.3.2); 11, phosphoenolpyruvatecarboxykinase (4.1.1.49) or phosphoenolpyruvate carboxylase (4.1.1.32);12, malic enzyme (1.1.1.40 or 1.1.1.38); 13, pyruvate carboxylase or oxaloacetate decarboxylase(6.4.1.1); 14, pyruvate dehydrogenase (solid line; 1.2.4.1 and 2.3.1.12) andpyruvate ferredoxin oxidoreductase (dashed line; 1.2.7.1). M. A. Huynen, T. Dandekar and P. Bork ``Variation and evolution of thecitric acid cycle: a genomic approach'' Trends Microbiol, 7, 281-29 (1999)
The citric-acid cycle b) Individual species mightnot have a complete CAC. This diagram shows the genes for the CAC for each unicellularspecies for which a genome sequence has been published, together with the phylogeny ofthe species. The distance-based phylogeny was constructed using the fraction of genesshared betweengenomes as a similarity criterion. The major kingdoms of life are indicatedin red (Archaea), blue (Bacteria) and yellow (Eukarya). Question marks representreactions for which there is biochemical evidence in the species itself or in a relatedspecies but for which no genes could be found. Genes that lie in a single operon areshown in the same color.Genes were assumed to be located in a single operon whenthey were transcribed in the same direction and the stretches of non-coding DNA separatingthem were less than 50 nucleotides in length. M. A. Huynen, T. Dandekar and P. Bork ``Variation and evolution of thecitric acid cycle: a genomic approach'' Trends Microbiol, 7, 281-29 (1999)
Small-world networks A recent paper, Collective dynamics of "small-world" networks, by Duncan J. Watts and Steven H. Strogatz, which appeared in Nature volume 393, pp. 440-442 (4 June 1998), has attracted considerable attention. One can consider two extremes of networks: The first are regular networks, where "nearby" nodes have large numbers of interconnections, but "distant" nodes have few. The second are random networks, where the nodes are connected at random. Regular networks are highly clustered, i.e., there is a high density of connections between nearby nodes, but have long path lengths, i.e., to go from one distant node to another one must pass through many intermediate nodes. Random networks are highly un-clustered but have short path lengths. This is because the randomness makes it less likely that nearby nodes will have lots of connections, but introduces more links that connect one part of the network to another.
Regular and random networks random regular regular complete
Regular, small-world and random networks:Rewiring experiments (Watts and Strogatz, 1998) p is the probability that a randomly chosen connection will be randomly redirected elsewhere (i.e.,p=0 means nothing is changed, leaving the network regular; p=1 means every connection is changed and randomly reconnected, yielding complete randomness). For example, for p = .01, (so that only 1% of the edges in the graph have been randomly changed), the "clustering coefficient" is over 95% of what it would be for a regular graph, but the "characteristic path length" is less than 20% of what it would be for a regular graph.
Small-world and networks A small-world network can be generated from a regular one by randomly disconnecting a few points and randomly reconnecting them elsewhere. Another way to think of a small world network is that some so-called 'shortcut' links are added to a regular network as shown here: The added links are shortcuts because they allow travel from node (a) to node (b), to occur in only 3 steps, instead of 5 without the shortcuts.
Small-world networks • Network characterisation: • L = characteristic path length • C = clustering coefficient • A small-world network is much more highly clustered than an equally sparse random graph (C >> Crandom), and its characteristic path length L is close to the theoretical minimum shown by a random graph (L ~ Lrandom). • The reason a graph can have small L despite being highly clustered is that a few nodes connecting distant clusters are sufficient to lower L. • Because C changes little as small-worldliness develops, it follows that small-worldliness is a global graph property that cannot be found by studying local graph properties.
Small-world networks A network or order (0<p<1 as in earlier slides) can be characterized by the average shortest length L(p) between any two points, and a clustering coefficient C(p) that measures the cliquishness of a typical neighbourhood (a local property). These can be calculated from mathematical simulations and yield the following behavior (Watts and Strogatz):
Small-world networks Part of the reason for the interest in the results of Watts and Strogatz is that small-world networks seem to be good models for a wide variety of physical situations. They showed that the power grid for the western U.S. (nodes are power stations, and there is an edge joining two nodes if the power stations are joined by high-voltage transmission lines), the neural network of a nematode worm (nodes are neurons and there is an edge joining two nodes if the neurons are joined by a synapse or gap junction), and the Internet Movie Database (nodes are actors and there is an edge joining two nodes if the actors have appeared in the same movie) all have the characteristics (high clustering coefficient but low characteristic path length) of small-world networks. Intuitively, one can see why small-world networks might provide a good model for a number of situations. For example, people tend to form tight clusters of friends and colleagues (a regular network), but then one person might move from New York to Los Angeles, say, introducing a random edge. The results of Watts and Strogatz then provide an explanation for the empirically observed phenomenon that there often seem to be surprisingly short connections between unrelated people (e.g., you meet a complete stranger on an airplane and soon discover that your sister's best friend went to college with his boss's wife).
Small world example: metabolism. • Wagner and Fell (2001) modeled the known reactions of 287 substrates that represent the central routes of energy metabolism and small-molecule building block synthesis in E. coli. This included metabolic sub-pathways such as: • glycolysis • pentose phosphate and Entner-Doudoro pathways • glycogen metabolism • acetate production • glyoxalate and anaplerotic reactions • tricarboxylic acid cycle • oxidative phosphorylation • amino acid and polyamine biosynthesis • nucleotide and nucleoside biosynthesis • folate synthesis and 1-carbon metabolism • glycerol 3-phosphate and membrane lipids • riboflavin • coenzyme A • NAD(P) • porphyrins, haem and sirohaem • lipopolysaccharides and murein • pyrophosphate metabolism • transport reactions • glycerol 3-phosphateproduction • isoprenoid biosynthesis and quinone biosynthesis • These sub-pathways form a network because some compounds are part of more than one pathway and because most of them include common components such as ATP and NADP. • Thegraphs on the left show that considering either reactants or substrates, the clustering coefficient C>>Crandom, and the length coefficient L is near that of Lrandom, characteristics of a small world system. random Wagner A, Fell D (2001) The small world inside large metabolic networks. Proc. R. Soc. London Ser. B 268, 1803-1810.
Scale-free Networks Using a Web crawler, physicist Albert-Laszlo Barabasi and his colleagues at the University of Notre Dame in Indiana in 1998 mapped the connectedness of the Web. They were surprised to find that the structure of the Web didn't conform to the then-accepted model of random connectivity. Instead, their experiment yielded a connectivity map that they christened "scale-free." • Often small-world networks are also scale-free. • Ina scale-free network the characteristic clustering is maintained even as the networks themselves grow arbitrarily large.
Scale-free Networks • In any real network some nodes are more highly connected than others. • P(k) is the proportion of nodes that have k-links. • For large, random graphs only a few nodes have a very small k and only a few have a very large k, leading to a bell-shaped Poisson distribution: Scale-free networks fall off more slowly and are more highly skewed than random ones due to the combination of small-world local highly connected neighborhoods and more 'shortcuts' than would be expected by chance. Scale-free networks are governed by a power law of the form: P(k) ~ k-
Scale-free Networks Because of the P(k) ~ k-power law relationship, a log-log plot of P(k) versus k gives a straight line of slope - : Some networks, especially small-world networks of modest size do not follow a power law, but are exponential. This point can be significant when trying to understand the rules that underlie the network.
Comparing Random and Scale-Free DistributionIn the random network (right), the five nodes with the most links (in red) are connected to only 27% of all nodes (green). In the scale-free network (left), the five most connected nodes (red), often called hubs, are connected to 60% of all nodes (green).
Scale-free Networks Before discovering scale-free networks, Barabasi and his team had been doing work that modeled surfaces in terms of fractals, which are also scale-free. Their discoveries about networks have been found to have implications well beyond the Internet; the notion of scale-free networks has turned the study of a number of fields upside down. Scale-free networks have been used to explain behaviors as diverse as those of power grids, the stock market and cancerous cells, as well as the dispersal of sexually transmitted diseases.
Scale-free Networks Put simply, the nodes of a scale-free network aren't randomly or evenly connected. Scale-free networks include many "very connected" nodes, hubs of connectivity that shape the way the network operates. The ratio of very connected nodes to the number of nodes in the rest of the network remains constant as the network changes in size. In contrast, random connectivity distributions—the kinds of models used to study networks like the Internet before Barabasi and his team made their observation—predicted that there would be no well-connected nodes, or that there would be so few that they would be statistically insignificant. Although not all nodes in that kind of network would be connected to the same degree, most would have a number of connections hovering around a small, average value. Also, as a randomly distributed network grows, the relative number of very connected nodes decreases.
Scale-free Networks The ramifications of this difference between the two types of networks are significant, but it's worth pointing out that both scale-free and randomly distributed networks can be what are called "small world" networks. That means it doesn't take many hops to get from one node to another—the science behind the notion that there are only six degrees of separation between any two people in the world. So, in both scale-free and randomly distributed networks, with or without very connected nodes, it may not take many hops for a node to make a connection with another node. There's a good chance, though, that in a scale-free network, many transactions would be funneled through one of the well-connected hub nodes - one like Google’s Web portal. Because of these differences, the two types of networks behave differently as they break down. The connectedness of a randomly distributed network decays steadily as nodes fail, slowly breaking into smaller, separate domains that are unable to communicate.
Scale-free Networks Resists Random Failure Scale-free networks, on the other hand, may show almost no degradation as random nodes fail. With their very connected nodes, which are statistically unlikely to fail under random conditions, connectivity in the network is maintained. It takes quite a lot of random failure before the hubs are wiped out, and only then does the network stop working. (Of course, there's always the possibility that the very connected nodes would be the first to go.) In a targeted attack, in which failures aren't random but are the result of mischief, or worse, directed at hubs, the scale-free network fails catastrophically. Take out the very connected nodes, and the whole network stops functioning. In these days of concern about cyber attacks on the critical infrastructure, whether the nodes on the network in question are randomly distributed or are scale-free makes a big difference.
Scale-free Networks Epidemiologists are also pondering the significance of scale-free connectivity. Until now, it has been accepted that stopping sexually transmitted diseases requires reaching or immunizing a large proportion of the population; most contacts will be safe, and the disease will no longer spread. But if societies of people include the very connected individuals of scale-free networks—individuals who have sex lives that are quantitatively different from those of their peers—then health offensives will fail unless they target these individuals. These individuals will propagate the disease no matter how many of their more subdued neighbors are immunized. Now consider the following: Geographic connectivity of Internet nodes is scale-free, the number of links on Web pages is scale-free, Web users belong to interest groups that are connected in a scale-free way, and e-mails propagate in a scale-free way. Barabasi's model of the Internet tells us that stopping a computer virus from spreading requires that we focus on protecting the hubs.
Scale Free Network •Hubs, highly connected nodes, bring together different parts of the network • Rubustness: Removing random nodes have little effect • Low attack resistance: Removing a hub is lethal (PPI: centrality-lethality rule). Random Network • No hubs • Low robustness • Low attack resistance
14-3-3 subtypes (paralogs) Schematic representation of co-immunoprecipitation studies performed with anti- MARK (microtubule affinity-regulating kinase) antibodies. The strength of the interactions is indicated by the thickness of the arrows (after (2) .
Preferential attachment • Hub protein characteristics: • Multiple binding sites • Promiscuous binding • Non-specific binding …connect preferentially to a hub
Hub proteins in yeast “[..] network analysis suggests that the centrality-lethality rule is unrelated to the network architecture, but is explained by the simple fact that hubs have large numbers of PPIs, therefore high probabilities of engaging in essential PPIs” He X, Zhang J (2006) Why do hubs tend to be essential in protein networks? PLoS Genet 2(6):e88
Network motifs • Different Motifs indifferent processes • • More interconnectedmotifs are moreconserved
Network Dynamics • Party hubs: always the samepartners (same time and space) • •Date hubs: different partners indifferent conditions (different time and/or space) • • Difference is important for inter-processcommunication Date hubs: large binding surfaces / Party hubs: small bindingsurfaces
A network example from Meta-genomics Ecogenomics – soil ecosystems A virtual network where species are nodes and (groups of) chemical compounds are exchanged between the nodes
Preferential attachment in biodegradation networks New degradable compounds are observed to attach preferentially to hubs close to (or in) the Central Metabolism Valencia and co-workers
The “Matchmaker” 14-3-3 family • Massively interacting protein family (the PPI champions) by means of various binding modes • Involved in many essential cell processes • Occurs throughout kingdom of life • Various numbers of isoforms in different organisms (7 in human)
14-3-3 network (hub?) promotion by binding and bringing together two different proteins