Decoding Genetics: From Genes to Proteins Blueprint
210 likes | 317 Views
Explore the journey from genes to proteins, starting with Garrod's hypothesis on genetic mutations. Learn how x-rays influenced genetic research, discover the role of enzymes in gene expression, and delve into the DNA blueprint for amino acids. Unravel the process of transcription and translation, understand the genetic code, and explore gene mutations' impact on protein production.

Decoding Genetics: From Genes to Proteins Blueprint
E N D
Presentation Transcript
Even before DNA was shown to be the genetic material a human geneticist had hypothesized & demonstrated how the blueprint would be organized. English physician A. Garrod proposed that if: Substrate A Product B Product C Product D Then a mutation leads to: gene 1 gene 2 gene 3 enzyme 1 enzyme 2 enzyme 3 gene 1 gene 2 gene 3 enzyme 1 enzyme 2 enzyme 3 Substrate A Product B Product C Product D
Even before DNA was shown to be the genetic material a human geneticist had hypothesized how the blueprint would be organized. English physician A. Garrod proposed that if: Substrate A Product B Product C Product D Then a mutation leads to: gene 1 gene 2 gene 3 enzyme 1 enzyme 2 enzyme 3 gene 1 gene 2 gene 3 enzyme 1 enzyme 2 enzyme 3 Substrate A Product B Product C Product D
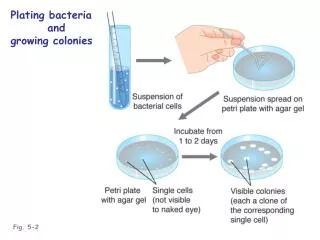
Early 1940's - Beadle & Tatum try to test Garrods idea They knew that x-rays could mutate the genetic blueprint they exposed bread mold to x-rays & looked for mutants that were unable to grow in minimal media Growth:Wild-typecells growingand dividing No growth:Mutant cellscannot growand divide Minimal medium If the hypothesis is correct then they should be able to rescue non-growers by adding in the missing enzymes product ….
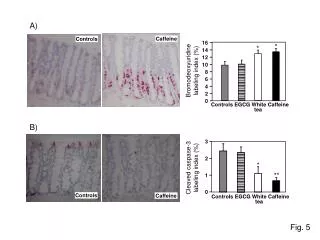
Beadle & Tatum found mutated mold strains that couldn't make arginine 'proved' the one gene-one enyzme hypothesis Fig 17.2
"knocking out" individual genes remains an important research tool …. determining cell fate & see thru animals tracking movement of fluorescent cells tracking metastatic melanoma movement Tumour
Fruit flies (Drosophila melanogaster) are another model organism that are used to study mutations in genes - especially for development
Early 1950's - still wasn't clear what proteins/enzymes were made of. In 1953 Frederick Sanger shows that core element of a protein is a sequence of amino acids. (many proteins are conjugated to other molecules e.g. glycoproteins) Therefore DNA sequence is the blueprint for amino acid sequence So a gene is just a segment of DNA that acts as a blueprint for amino acids!? Or is there more to a gene?
Defining a gene and it’s properties. A gene is all the information contained within a sequence of DNA that is required to make a single complete polypeptide. (not 100 % true) Not all the information is for the design of the protein: additional information will direct how & when the polypeptide is made some information may direct where in the cell the polypeptide is taken So the gene contains both instructional information & design information
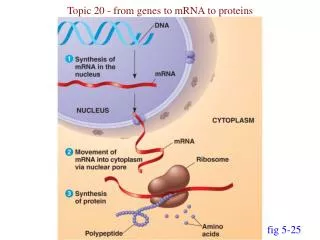
To get the information within a gene conveyed into a polypeptide …. First pare down genetic information to a minimum - called messenger RNA or just mRNA The process of making the mRNA from the gene is called transcription The information within the mRNA is then translated, from nucleic acid to amino acid by a process is called translation.
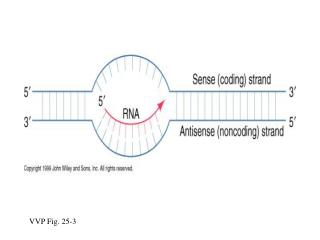
Going from gene to protein involves two distinct steps: DNAtemplatestrand DNA 5 3 molecule A A A A A T C C C C G G T T T T A G G G G C T C Figure 17.4 Gene 1 3 5 TRANSCRIPTION Gene 2 U G G U U U G G C U C A 5 3 mRNA ? ? TRANSLATION Gly Phe Trp Protein Ser Gene 3 Amino acid With 20 amino acids & only 4 nucleotide types how do you convert genetic code to protein?
A code based on three nucleotides gives 64 possible codons (43) – more than enough for 20 AAs –and with some room for errors DNAtemplatestrand DNA 5 3 molecule A A A A A Figure 17.4 T C C C C G G T T T T A G G G G C T C Gene 1 3 5 TRANSCRIPTION Gene 2 U G G U U U G G C U C A 5 3 mRNA Codon TRANSLATION Gly Phe Trp Protein Ser Gene 3 Amino acid
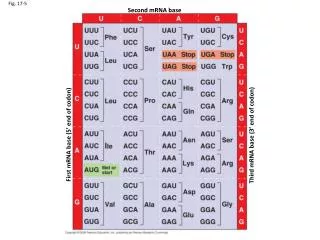
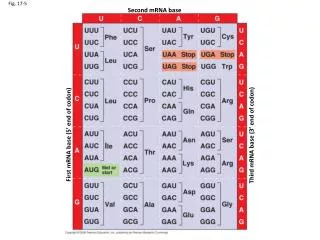
Translating from nucleotide codons to amino acids The genetic code is said to be degenerate (has ‘room for error’) fig 17.5
Almost all organisms use the same genetic code The genes encoding the 'glow' proteins of fireflies, jelly fish, and other invertebrates were used to make these pet-store Glo-fish Many bacteria would also be able to read these 'glow' genes. mitochondria & some protists use a somewhat distinct genetic code
Given the degenerate nature of the code a single nucleotide mutation may or may not alter the encoded protein Fig 17.2
Gene mutations can alter single codons or the whole reading frame. Two types: Base pair substitution A new codon is created & possibly a new amino acid encoded Outcomes: Silent mutation - no change to AA encoded Mis-sense - codon change causes AA change Non-sense - codon changes to stop codon
e.g. mutating the Cysteine codon from UGU to UGC would not change the encoded AA – it’s a silent mutation UGU to UGA would lead to a stop codon – a non-sense mutation & UGU to UGG would change the AA to Tryptophan – a mis-sense mutation fig 17.5
Gene mutations can alter single codons or the whole reading frame. Two types: Base pair substitution 2) Removing or adding in base pairs will alter the reading frame, unless its in multiples of 3 bases most codons will change to missense/nonsense codons
Because the codons are read as a continuous sequence the same RNA sequence could be read in different ways - 3 reading frames CCACCUCCCCCGAUUUACCCAcan be read: CCA CCU CCC CCG AUA AAG CCA P P P P I K P amino acid abbreviations But if the first cytosine was deleted you’d ‘frameshift’ how you read to: C CAC CUC CCC CGA UAA AGC CA H L P R * S *stop codon - end of open reading frame
Two final questions…. 1) Are all genes translated into amino acid sequence? 2) Is all the DNA a part of a gene? i.e. is a strand of DNA just one gene after another without interruption?
1) The information contained in a gene may not always be used to make a polypeptide. Sometimes the functional end product is the RNA. There are examples of functional RNAs used in translation There are even enzymes made of RNA called “Ribozymes” 2) In eukaryotes the genes are separated from one another with non-coding DNA called inter-genic DNA