[S1]
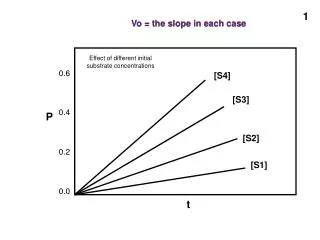
Vo = the slope in each case. Vo = the slope in each case. Effect of different initial substrate concentrations. 0.6. [S4]. [S3]. 0.4. P. [S2]. 0.2. [S1]. 0.0. t. No enzyme. Considering Vo as a function of [S] (which wil be our usual useful consideration). Now, with an enzyme:.

[S1]
E N D
Presentation Transcript
Vo = the slope in each case Vo = the slope in each case Effect of different initial substrate concentrations 0.6 [S4] [S3] 0.4 P [S2] 0.2 [S1] 0.0 t
No enzyme Considering Vo as a function of [S] (which wil be our usual useful consideration)
Now, with an enzyme: We can ignore the rate of the non-catalyzed reaction
Enzyme catalyzed kinetics (as opposed to simple chemical kinetics) Vo independent of [S] Vo proportional to [S]
Michaelis and Menten mechanism for the action of enzymes (1913)
Michaelis-Menten mechanism • Assumption 1. E + S <--> ES: this is how enzymes work, via a complex • Assumption 2. Reaction 4 is negligible, when considering INITIAL velocities (Vo, not V). • Assumption 3. The ES complex is in a STEADY-STATE, with its concentration unchanged with time during this period of initial rates. (Steady state is not an equilibrium condition, it means that a compound is being added at the same rate as it is being lost, so that its concentration remains constant.) X
System is at “steady state” Constant level Plenty of flow Steady state is not the same as equilibrium System is at equilibrium Constant level No net flow
E + S ES E + P System is at “steady state” Constant level Plenty of flow
k3 [Eo] [S] Vo = Km + [S] Michaelis-Menten Equation(s) See handout at your leisure for the derivation (algebra) k3[Eo][S] Vo = [(k2+k3)/k1] +[S] If we let Km = (k2+k3)/k1, just gathering 3 constants into one, then:
Rate is proportional to the amount of enzyme Otherwise, the rate is dependent only on S k3 [Eo] [S] Vo = Km+ [S] All the k‘s are constants for a particular enzyme At high S (compared to Km), Rate is constant Vo = k3Eo At low S (compared to Km), rate is proportional to S: Vo ~ k3Eo[S]/Km
k3 [Eo] [S] Vo = Km + [S] At high S, Vo here = Vmax = k3Eo So the Michaelis-Menten equation can be written: Vmax [S] Vo = Simplest form Km + [S]
Now, Vmax = k3Eo So: k3 = Vmax/Eo = the maximum (dP/dt)/Eo, = the maximum (-dS/dt)/Eo k3 = the TURNOVER NUMBER • the maximum number of moles of substrate converted to product per mole of enzyme per second; • max. molecules of substrate converted to product per molecule of enzyme per second • Turnover number then is: a measure of the enzyme's catalytic power.
Some turnover numbers • Succinic dehydrogenase: 19 (below average) • Most enzymes: 100 -1000 • The winner: Carbonic anhydrase (CO2 +H20 H2CO3) 600,000 That’s 600,000 molecules of substrate per molecule of enzyme per second. Picture it! You can’t.
Consider a Vo that is 50% of Vmax Km ? Vmax/2 is achieved at a [S] that is numerically equal to Km So Km is numerically equal to the concentration of substrate required to drive the reaction at ½ the maximal velocity Try it: Set Vo = ½ Vmax and solve for S.
k2 ES E + S k1 Another view of Km: Consider the reverse of this reaction (the DISsociation of the ES complex): The equilibrium constant for this dissociation reaction is: Kd = [E][S] / [ES] = k2/k1 (Note: This equation has been corrected from the original ) (It’s the forward rate constant divided by the backward rate constant. See the Web lecture if you want to see this relationship derived)
k2 ES E + S k1 { Consider in reverse Kd = k2/k1 Km = (k2+k3)/k1 IF k3 << k2, then: Km ~ k2/k1 But Kd = k2/k1 (from last graphic) so Km ~ Kd for the dissociation reaction (and 1/Km = ~ the association constant) So: the lower the Km, the more poorly it dissociates. That is, the more TIGHTLY it is held by the enzyme And the greater the Km, the more readily the substrate dissociates, so the enzyme is binding itpoorly
Km ranges • 10-6M is good • 10-4M is mediocre • 10-3M is fairly poor So Km and k3 quantitatively characterize how an enzyme does the job as a catalyst
Apparent Km increases - Inhibitor looks like the substrate and like the substrate binds to the substrate binding site
Competitive inhibitor can be swamped out at high substrate concentrations Handout 5-3b
Apparent Km increases - Inhibitor looks like the substrate and like the substrate binds to the substrate binding site
- Inhibition eventually swamped out as S is increased
½ Vmax w/o inhibitor ½ Vmax withyet more inhibitor Km remains unchanged. Vmax decreases.
Substrate Non-competitive inhibitor Example: Hg ions (mercury) binding to –SH groups in the active site
Hg++ --CH2-SH --CH2-SH Non-competitive inhibitor example Substrate still bonds OK But an essential participant in the reaction is blocked (here, by mercury binding a cysteine sulfhydryl)
Allosteric inhibition Inhibitor binding site + Active Active Inactive = allosteric inhibitor = substrate Allosteric inhibitor binds to a different site than the substrate, So it need bear no resemblance to the substrate The apparent Km OR the apparent Vmax or both may be affected.
Allosteric inhibitors are used by the cell for feedback inhibition of metabolic pathways Feedback inhibition of enzyme activity, or End product inhibition End product End product End product First committed step is usually inhibited
protein Thr deaminaseglucose ...... --> --> threonine -----------------> alpha-ketobutyric acid protein A Substrate B C isoleucine (and no other aa) Allosteric inhibitor Also here: Feedback inhibitor (is dissimilar from substrate)
20 minutes !, in a rich medium 60 minutes, in a minimal medium
Direction of reactions in metabolism
} Energy difference Free determines the direction of a chemical reagion
For the model reaction A + B C + D, written in the left-to-right direction indicated: Consider the quantity called the change in free energy associated with a chemical reaction, or: ΔG Such that: • IF ΔG IS <0:THEN A AND BWILL TEND TO PRODUCE C AND D(i.e., tends to go to the right). • IF ΔG IS >0:THEN C AND D WILL TEND TO PRODUCE A AND B.(i.e., tends to go to the left) • IF ΔG IS = 0:THEN THE REACTION WILL BE AT EQUILIBRIUM: NOT TENDING TO GO IN EITHER DIRECTION IN A NET WAY.
ΔG = ΔGo+ RTln([C][D]/[A][B]) • where A, B, C and D are the concentrations of the reactants and the products AT THE MOMENT BEING CONSIDERED.(i.e., these A, B, C, D’s are notthe equilibrium concentrations) • R = UNIVERSAL GAS CONSTANT = 1.98 CAL / DEG K MOLE • T = ABSOLUTE TEMP. (0oC = 273oK; Room temp = 25oC = 298o K) • ln = NATURAL LOG • Δ Go = a CONSTANT: a quantity related to the INTRINSIC properties of A, B, C, and D
Josiah Willard Gibbs(1839 - 1903) Also abbreviated form:ΔG = ΔGo+ RTlnQ Where Q = ([C][D]/[A][B]) Qualitative term Quantitative term
ΔGo STANDARD FREE ENERGY CHANGE of a reaction. If all the reactants and all the products are present at 1 unit concentration, then: ΔG = ΔGo + RTln(Q) = ΔGo + RTln([1][1] / [1][1]) ΔG = ΔGo + RTln(Q) = ΔGo + RTln(1) or ΔG = ΔGo +RT x 0, or ΔG = ΔGo, when all components are at 1M ….. a special case (when all components are at 1M)
So ΔG and ΔGo are quite different, and not to be confused with each other. ΔGo allows us to compare all reactions under the same standard reaction conditions that we all agree to, independent of concentrations. So it allows a comparison of the stabilities of the bonds in the reactants vs. the products. It is useful. AND, It is easily measured.
[C]eq [D]eq [A]eq [B]eq Because, • at equilibrium, ΔG= ΔGo + RTln(Q) = 0 and at equilibrium Q = Keq = (a second special case). • So: at equilibrium,ΔG= ΔGo + RTln(Keq) = 0 • And so: ΔGo = - RTln(Keq) • So just measure the Keq, • Plug in R and T • Get: ΔGo, the standard free energy change
E.g., let’s say for the reaction A + B C + D, Keqhappens to be: = 2.5 x 10-3 [C]eq[D]eq [A]eq[B]eq Then ΔGo = -RTlnKeq = - 2 x 300 x ln(2.5 x 10-3) = -600 x -6 = +3600 3600 cal/mole (If we use R=2 we are dealing with calories) Or: 3.6 kcal/mole 3.6 kcal/mole ABSORBED (positive number) So energy is required for the reaction in the left-to-right direction And indeed, very little product accumulates at equilibrium (Keq = 0.0025)
Note: IfΔGo = +3.6 for the reaction A + B < --- >C + D ThenΔGo = -3.6 for the reaction C + D <--- >A + B (Reverse the reaction: switch the sign) And: For reactions of more than simple 1 to 1 stoichiometries: aA + bB <--> cC + dD, ΔG = ΔGo + RT ln [C]c[D]d [A]a[B]b
Some exceptions to the 1M standard condition: Exception #1: • 1) Water: 55 M (pure water) is considered “unit” concentration instead of 1MThe concentration of water rarely changes during the course of an aqueous reaction, since water is at such a high concentration. • So when calulating Go, instead of writing in “55” when water participates in a reaction (e.g., a hydrolysis) we write “1.” • This is not cheating; we are in charge of what is a “standard” condition, and we all agree to this: 55 M H20 is unit (“1”) concentration for the purpose of defining Go.
Exception #2: In the same way, Hydrogen ion concentration, [H+]: 10-7 M is taken as unit concentration, by biochemists. since pH7 is maintained in most parts of the cell despite a reaction that may produce acid or base. This definition of the standard free energy change requires the designation ΔGo’ However, I will not bother. But it should be understood we are always talking about ΔGo’ in this course.
Summary ΔG= Go + RTln(Q) This combination of one qualitative and one quantitative (driving) term tell the direction of a chemical reaction in any particular circumstance ΔGo = - RTln(Keq) TheΔGo for any reaction is a constant that can be looked up in a book.
ATP, a small molecule in the cell that helps in the transfer of energy from a place where it is generated to a place where it is needed. e e A-R
The hydrolysis of ATP: ATP + HOH ADP + Pi A= adenine (2 rings with N’s) R = ribose (5-carbon ring sugar) ADP AMP The Go of this reaction is about -7 kcal/mole. Energy is released in this reaction. This is an exergonic reaction Strongly to the right, towards hydrolysis, towards ADP
O O O || || || A-R-O-P-O-P-O- + -O-P-O- | | | O- O- O- O O O || || || A-R-O-P-O-P-O-P-O- + HOH | | | O- O- O- ATP Adenosine triphosphate ADP Adenosine diphosphate Pi Inorganic phosphate The Go of this reaction is about -7 kcal/mole. Energy is released in this reaction. This is an exergonic reaction Strongly to the right, towards hydrolysis, towards ADP
“High energy” bonds: • Gomore negative than ~ -7 kcal/mole is released upon hydrolysis • Designated with a squiggle (~): • ATP = A-P-P~P • Rationalized by the relief of electrical repulsion upon hydrolysis: ΔGo = -7 kcal/mole
AMP ΔGo = -7 kcal/mole ADP A-R-P~P~P Not a high energy bond ATP + HOH + ATPase ADP + Pi + heat Prob set 4 Keq ~ 100,000