Bacteriophage
Bacteriophage . Khansa Bukhari Maryum Zeb. History and Definition. Frederick Twort (1915) and Felix d'Herelle (1917) were the first to recognize bacteriophage. In the 1930s and after words virologists utilized these viruses as model systems to investigate many aspects of virology

Bacteriophage
E N D
Presentation Transcript
Bacteriophage KhansaBukhari MaryumZeb
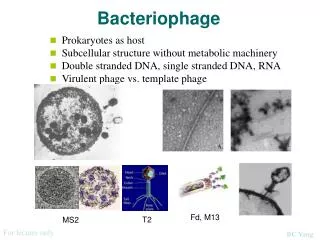
History and Definition • Frederick Twort (1915) and Felix d'Herelle (1917) were the first to recognize bacteriophage. • In the 1930s and after words virologists utilized these viruses as model systems to investigate many aspects of virology Viruses which infect bacteria are called bacteriophages (eaters of bacteria).
Classification • Phages are classified by the International Committee on Taxonomy of Viruses (ICTV) according to morphology and nucleic acid.
Bacteriophage M13 • Belongs to family Inoviridae • Filamentous • Circular single-stranded DNA • 6400 base pairs long • It infects F+ E. coli • It just slows growth of host cell • Virions leak out from the cell. • The virus is not very lytic.
Genome • The genome codes for a total of 10 genes (named using Roman numerals I - X) • Gene VIII codes for the major structural protein, forms a tubular array of approx. 2,700 identical subunits surrounding the viral genome • Gene III codes for the minor coat protein • Approximately five to eight copies of the gene III protein are located at the ends of the filamentous phage • Allows binding to bacterial "sex" pilus • Pilus is a bacterial surface structure of E. coli which harbor the "F factor" extrachromosomal element
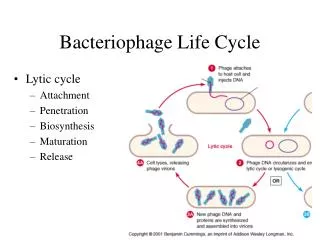
REPLICATION • Single strand genome ( '+' strand) attached to pilus enters host cell • Major coat protein (gene VIII) stripped off • Minor coat protein (gene III) remains attached • Host components convert single strand (+) genome to double stranded circular DNA (replicative form or "RF") • Transcription takes place by host RNA polymerase • Viral g2p protein nicks RF DNA strand at the origin of replication. • (+)strand replication occurs by rolling circle. • New (+)ssDNA genomes are converted into new RF molecules, and further transcription occurs. • When enough g5p protein is synthesized, conversion into RF dsDNA is inhibited, as neo-synthesized genomic ssDNA is covered with g5p. • g5p are replaced by g8p proteins to assemble the viral capsid. • New virions bud out from host cell. • Infected cells continue to divide and produces virion indefinitely.
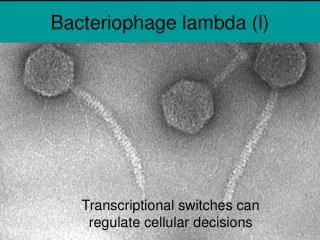
Classification • Virus classification • Group: Group I (dsDNA) • Order: Caudovirales • Family: Siphoviridae • Genus: λ-like viruses • Species: λ Phage
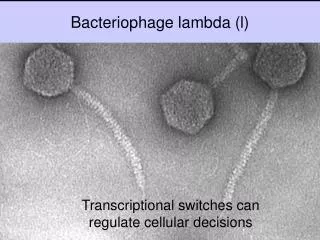
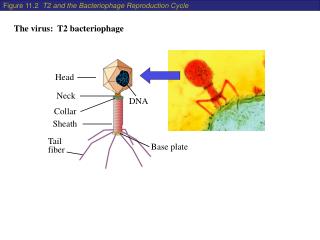
Bacteriophage lambda • Bacteriophage lambda (λ) was discovered by Joshua and Esther Lederberg. While mutagenizing strains E. coli using UV • Genome is 48,502 bpdsDNA • Linear dsDNA circularizes due to annealing between 3′ sticky cos sites. • The capsid has two parts: • Head, composed of B, C, Nu3, D, and E proteins. • Tail, composed of J and H proteins. • It is a temperate phage: being either lytic or lysogenic. • Chooses developmental pathway depending on nutrient availability
Classification • Group group III, dsRNA virus • Family Cystoviridae • Genus Cystovirus • Specie Pseudomonas phage phi6
Structure & Entry • Multi-segmented, Spherical enveloped virus of Pseudomonas syringae • The nucleocapsid NC, consisting of the five proteins P1 , P2, P4, P7 and P8 • Entry begins by viral attachment, followed by fusion with the host cell outer membrane • NC penetrates the cytoplasmic membrane via membrane invagination with the aid of protein P8. • steps releases a transcriptionally active polymerase complex called the core into the cytoplasm.
Genome • Genome consists of three segments: small (S), medium (M) and large (L). • Core is encoded by L segment. • P1 is the major structural protein • P4 is identified as nucleoside triphosphotase, which is necessary for +strand RNA packaging. • P7 stabilizes RNA packaging. • P2 is rdrp, carries out –strand synthesis and RNA transcription.
Replication • Pseudomenassynirgaepilibrings the virion into contact with the NC associated lytic enzyme locally digests the peptidoglycan. • Upon entry, viral polymerase is activated to produce early transcripts. • Translation of L transcripts produces the early proteins, which assemble to form PC. • Packaging of +s transcripts and synthesis of –s transcripts inside the PC
Transcription by these PC produces message for late gene synthesis • The NC surface proteins assemble on the PC and inactivated transcription. • The NC acquires membrane from the host plasma membrane. • The cell eventually lyses and liberates mature progeny particles.
Phage Therapy • Use of phages to treat bacterial infections in animals and humans. • treat bacterial dysentery, staphylococcal lung infections, surgical wound infections, • staphylococcal septicemia • Easier to develop new phage than a new antibiotic. • Replicate at the site of infection and are thus available where they are most needed • Phage-resistant bacteria remain susceptible to other phages having a similar target range
Phage Lysins as Antimicrobials • Use of phage endolysins as potential therapeutics instead of whole phage. • Phage endolysins, or lysins, are enzymes that damage the cell walls' integrity • phage lysinscomposed of at least two distinctly separate functional domains: • C-terminal cell-wall binding domain, which directs the enzyme to its target • N-terminal catalytic domain
Phage Display • 1. Insert a diverse group of genes into the phage genome. Each phage receives a different gene. • 2. Create a library of genetically modified phages which are all related but each of which have a different gene. • 3. Isolate a disease causing molecule, such as a receptor or an enzyme, and expose the phage library to the molecule. • 4. Some of the phages will bind to the disease causing molecule. Wash the phage library. All of the phages with proteins that didn't bind to the disease causeing molecule will be removed. • 5. Replicate the phages that remain so that they can be sequenced. • 6. Since we know where the new gene was inserted we can determine the amino acid sequence of the protein that bound to the disease
Applications of Phage Display • To select proteins, peptides, or antibodies with affinity to a molecule or protein of interest • To clone antibodies from unstable hybridoma. • To identify molecules that can be recognized and internalized by eukaryotic cells. • To identify epitopes, functional and accessible sites from antigens. • To study protein-protein interaction. • To design vaccines.
Use of Phage in Vaccination • Whole bacteriophage acts as efficient DNA vaccine delivery vehicles. • Phage-display vaccination. • Phage DNA vaccination. • Hybrid phage vaccine.