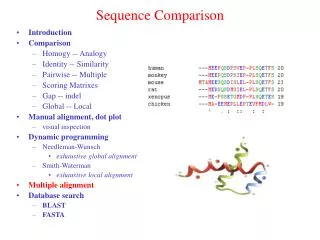
Dynamic Programming in Genome Sequence Comparison
Explore sequence alignment algorithms using dynamic programming for genetic analysis. Learn scoring methods and align sequences efficiently.

Dynamic Programming in Genome Sequence Comparison
E N D
Presentation Transcript
Sequence comparison: Dynamic programming Genome 559: Introduction to Statistical and Computational Genomics Prof. William Stafford Noble
One-minute responses • The pace was good until the end. One or two more minutes for each sample problem would have been good. • The pace worked well for me. I scrambled a bit to keep up on the computer but was able to do so. • I feel like a fish out of water, but the pace of the class was good. With this kind of stuff it’s really easy to feel stupid if you haven’t done it before, so I like the relaxed intro feeling. • I thought the pace and amount of material was fine. • Thought it covered the basics, didn’t move too fast. • I am very excited about this class. I enjoyed the slow and steady pacing because I am a beginner in programming. The sample problems are excellent. I wish there was time for more. • It seemed that the class was a bit too fast. It was hard to keep up. I was not sure what I was doing! Perhaps prior reading would help. • Thanks for moving so slowly and being so patient. Please keep it up. • I have no background in programming, and though I recognize that today’s class was very basic, I struggled some. I am not sure how I would have had an easier time – perhaps if more time had been taken explaining how each component (import, math, commas, etc.) had impact on function.
One-minute responses • It is still unclear to me why the “float” prefix is necessary. • Because otherwise Python will define the argument as a string, rather than a number. • I tried importing math and then using math.sum(arg1, arg2) because it seemed logical after the log example. • This was an error on my part – I should have introduced the “+” operator before that sample problem. • Is what we’re referring to as an “argument” anything that follows the program name on the command line? • Yes. • Is that generally what is meant by an argument, or does it have a broader meaning? • The broader meaning is anything that is given as input to a function; e.g., 10 is the argument to math.log(10). In general, Python functions take arguments in parentheses. (The print command is an exception to this rule.) • I like that the instructors walk around to check on things. • It might help to explain where we can look up packages to do various functions. • Your book describes the most common packages. Otherwise, you can search at python.org. • I think some simple terms could be defined like “terminal,” “system,” “argument.” • The programming exercises were useful, but if I didn’t already have a basic knowledge of Perl I would be confused by the terminology (operators, variables, objects, working directory, etc.). • This was my first time programming, so I was confused by terminology. After seeing the examples, I started to understand it better, though.
One-minute responses • One thing I may like is if we could have an appendix for the commands we learned that day. I imagine maybe the book has it? • All of the commands you learn are in the book. I encourage each of you to maintain a summary of what we’ve learned thus far. • Could you give supplementary problems to work on? • Yes, I will try to do this. • It was helpful having a printed copy of the lab task. • Can you print quotation marks? Are there other symbols that are off-bounds to print? • You can print anything. Unusual characters need to be preceded by a backslash: print "\"" • It was a little unclear at first what was the relevance of the sys.argv command. • I probably missed it but the only thing I could think of was more advanced warning on the text. • It was hard to follow directions without having read for myself to put into proper context.
Sequence comparison overview • Problem: Find the “best” alignment between a query sequence and a target sequence. • To solve this problem, we need • ___________________, and • ___________________. • The alignment score is calculated using • ___________________, and • ___________________. • The algorithm for finding the best alignment is __________________.
Sequence comparison overview • Problem: Find the “best” alignment between a query sequence and a target sequence. • To solve this problem, we need • a method for scoring alignments, and • an algorithm for finding the alignment with the best score. • The alignment score is calculated using • a substitution matrix, and • gap penalties. • The algorithm for finding the best alignment is dynamic programming.
Review • What does the 62 in BLOSUM62 mean? • Why does leucine and isoleucine get a BLOSUM62 score of 2, whereas leucine and aspartic acid get a score of -4? • What is the difference between a gap open and a gap extension penalty?
Review • What does the 62 in BLOSUM62 mean? • The sequences in the alignment used to generate the matrix share 62% pairwise sequence identity. • Why does leucine and isoleucine get a BLOSUM62 score of 2, whereas leucine and aspartic acid get a score of -4? • Leucine and isoleucine are biochemically very similar; consequently, a substitution of one for the other is much more likely to occur. • What is the difference between a gap open and a gap extension penalty? • When we assign a score to an observed gap in an alignment, we charge a larger penalty for the first gapped position than for subsequent positions in the same gap.
A simple alignment problem. • Problem: find the best pairwise alignment of GAATC and CATAC. • Use a linear gap penalty of -4. • Use the following substitution matrix:
How many possibilities? GAATC CATAC GAAT-C C-ATAC -GAAT-C C-A-TAC • How many different alignments of two sequences of length N exist? GAATC- CA-TAC GAAT-C CA-TAC GA-ATC CATA-C
How many possibilities? GAATC CATAC GAAT-C C-ATAC -GAAT-C C-A-TAC • How many different alignments of two sequences of length n exist? GAATC- CA-TAC GAAT-C CA-TAC GA-ATC CATA-C Too many to enumerate!
-G- CAT DP matrix The value in position (i,j) is the score of the best alignment of the first i positions of the first sequence versus the first j positions of the second sequence. -8
-G-A CAT- DP matrix Moving horizontally in the matrix introduces a gap in the sequence along the left edge.
-G-- CATA DP matrix Moving vertically in the matrix introduces a gap in the sequence along the top edge.
G - Introducing a gap
- C DP matrix
G C DP matrix
----- CATAC DP matrix
-G CA G- CA --G CA- DP matrix -4 -9 -12 0 -4 -4
DP matrix What is the alignment associated with this entry?
DP matrix -G-A CATA
DP matrix Find the optimal alignment, and its score.
GA-ATC CATA-C DP matrix
GAAT-C CA-TAC DP matrix
GAAT-C C-ATAC DP matrix
GAAT-C -CATAC DP matrix
Multiple solutions • When a program returns a sequence alignment, it may not be the only best alignment. GA-ATC CATA-C GAAT-C CA-TAC GAAT-C C-ATAC GAAT-C -CATAC
DP in equation form • Align sequence x and y. • F is the DP matrix; s is the substitution matrix; d is the linear gap penalty.
Dynamic programming • Yes, it’s a weird name. • DP is closely related to recursion and to mathematical induction. • We can prove that the resulting score is optimal.
Summary • Scoring a pairwise alignment requires a substition matrix and gap penalties. • Dynamic programming is an efficient algorithm for finding the optimal alignment. • Entry (i,j) in the DP matrix stores the score of the best-scoring alignment up to those positions. • DP iteratively fills in the matrix using a simple mathematical rule.
Reading • Nicholas review from Biotechniques, 2000.
Problem: find the best pairwise alignment of GAATC and CATAC. d = -4