Proteomics:
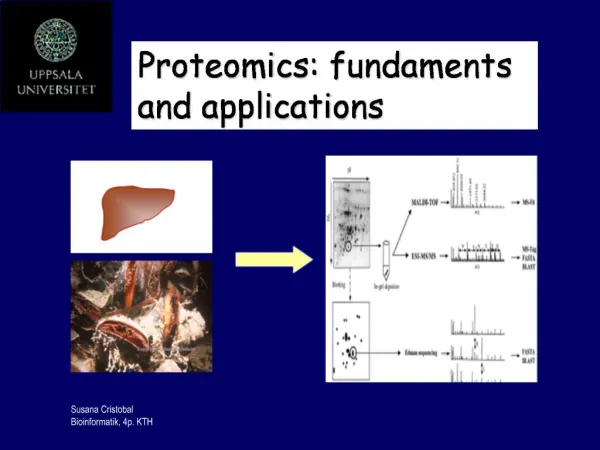
Proteomics:. Its Function and Methods. Ryan Victor. Proteomics. What Is It? What’s Being Studied? How Is It Done? Two-Dimensional Electrophoresis. What is It?. Proteomics is the study of proteins that are generated from the genetic code of an organism.

Proteomics:
E N D
Presentation Transcript
Proteomics: Its Function and Methods Ryan Victor
Proteomics What Is It? What’s Being Studied? How Is It Done? Two-Dimensional Electrophoresis
What is It? Proteomics is the study of proteins that are generated from the genetic code of an organism. Proteomics differs from genomics in that the chromosomes for a genome are consistent throughout a multicellular organism, protein output varies from cell to cell based on the cells function. While the genetic code for humans has been completed through the human genome project, proteomics will be a much longer, more intensive and time consuming study.
What is It? Initial attempts to study proteins used mRNA as the determining factor for protein level output. However, post translational modifications make the method invalid. The study of proteomics is forced to focus on very stringent conditions in a cell in order for the results to be valid. The cell must change its protein output based on it’s level of stress and metabolic needs. The results of a proteomics study can really only be generalized to the conditions that the cell faced during the study.
What’s Being Studied Phosphorylation: often is used to activate a protein, usually on a serine or threonine residue. Activation occurs as the phosphorylation is used to signal other proteins and thus permit binding between them. Ubiquination: often is a signal for a protein to be degraded. Once a protein has ubiquitin attached to it, it can be labeled for destruction by proteasome.
How to Study It Antibody Method: • Antibodies are adding to the protein mixture • Antibodies bind to proteins that have modified • Proteins of interest can be separated based on the modification.
2 Dimensional Protein Gels -What’s the Difference? -How’s It Done? -What Can We Do With It Fluorescently labeled proteins in difference gel electrophoresis
The Difference Conventional protein gels only run in one dimension, as the name implies, 2D gels run in 2 dimensions. 2nd running dimension allows separation based on a 2nd characteristic. Common separation characteristics are isoelectric point in the first dimension, followed by mass/size separation in the 2nd dimension.
How It’s Done We’ll look at isoelectric focusing as an example: Proteins are fed into a gel medium that has a pH gradient to it. pH gradient is formed by adding polyampholytes to the gel medium or using a gradient gel. Polymapholytes are much like amino acids in that they are zwitterionic.
How It’s Done Proteins each have their own isoelectic point that comes about as an average of their amino acid isoelectic point. Proteins are charged at different pHs. When they reach the region of the gel that matches their isoelectric point, they stop moving. Proteins can be removed at this point for further analysis if desired. Proteins can also be subjected to further analysis within the gel.
How It’s Done Now that the proteins have been separated by isoelectic point, they can be analyzed based on their mass. Proteins are separated by mass using Sodium Dodecyl Sulfate. SDS acts as a detergent to uncoil the protein and give it a negative charge, since the proteins have zero charge after the isoelectic focusing. The proteins migrate by applying an electric field 90 degrees from where it was located for isoelectric focusing. Proteins have now migrated in 2 different dimensions from their starting point, giving a “3D” gel.
How It’s Done Now that the proteins have migrated to their new positions based on isoelectric point and mass, they can be analyzed. Gel can be stained using coomassie or silver. Silver binds to the cysteine groups in the protein, and leaves a dark stain on the gel after development. Coomassie binds to arginine, histidine, and aromatic amino acids. It can also be used to replace SDS to give the negative charge to the protein.
Application Comparing the protein output between two different organism samples Comparing output between cells within a specific organism Determining whether an organism lacks a specific protein output Associating protein output with a specific disease, and thus aiding in drug development
Problems Two dimensional gels must still be read, which has its own variety of difficulties - Spots can overlap - Gel may not be reproducible - Spots may not properly visualize and be too weak to see. Duplicate gels overlaid upon each other in the Delta 2D program Scanners and computer programs are used to read the gels. They are only as good as the scanners acuity and the programs ability to discern spots on the scan.
Difference Gel Electrophoresis Fluorescent dye is added to the proteins, then the gel is analyzed using a laser to excite the fluorescent dyes. The abundance of proteins can be viewed from different samples to give an idea of the difference between them. Eliminates the difficulty of comparing different gels.
Summary Proteomics allows researchers to study the actual output of the cells rather than just the DNA blueprints. Proteomics can be used to develop effective treatments for diseases. 2-Dimensional Electrophoresis is an effective method for visualizing the results of a proteomics experiment. 2D gels are not without their difficulties.
References Unlu, M., Morgan, M., Minden J. (1997). Difference Gel Electrophoresis. Electrophoresis 11, 2071-7 King, M. (2009, November 2nd). Protein Modifications. Retrieved March 5th 2009 from The Medical Biochemistry Page website, http://themedicalbiochemistrypage.org/protein-modifications.html Anderson, N. (2005). Proteome and Proteomics. Electrophoresis 19, 1853-1861 Maiman Institute for Proteome Research. Two-Dimensional Gel Electrophoresis. Retrieved March 5th 2009 from the Maimane Institute for Proteome Research at Tel Aviv website, http://www.tau.ac.il/lifesci/units/proteomics/2dimgel.html Berth, M. et al. (2007) The State of the Art in Analysis of Two-Dimensional Gel Electrophoresis Images. Applied Microbiology Biotechnology 76, 1223-43