Flow of genetic information
470 likes | 1.28k Views
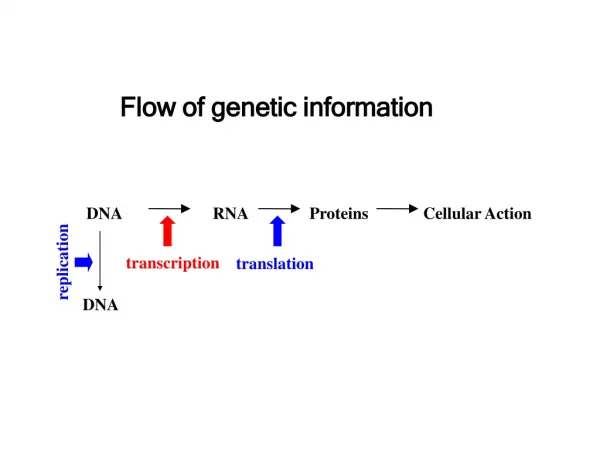
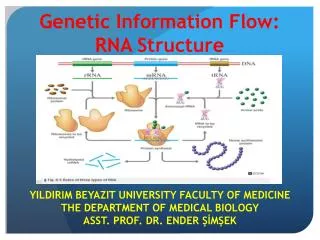
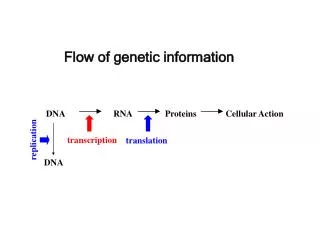
Flow of genetic information. DNA. RNA. Proteins. Cellular Action. replication. transcription. translation. DNA. RNA Types. mRNA = Messenger RNA; an RNA copy of the DNA sequence (gene) used as a template for protein synthesis

Flow of genetic information
E N D
Presentation Transcript
Flow of genetic information DNA RNA Proteins Cellular Action replication transcription translation DNA
RNA Types • mRNA = Messenger RNA; an RNA copy of the DNA sequence (gene) used as a template for protein synthesis • tRNA = Transfer RNA; a small RNA that is attached to an amino acid which can be added to a growing peptide chain • rRNA = Ribosomal RNA; component of ribosomes with catalytic and structural function • snRNA = Small nuclear RNA, involved in RNA splicing in • eukaryotes
2Pi RNA synthesis • RNA synthesis involves transcribing a specific portion of DNA strand • into RNA sequence • RNA polymerases sequentially add ribonucleotides to the 3’ end • of an RNA polymer using DNA strand as a template (5’ 3’ direction) • RNA Pol selects ribonucleotides complementary to the DNA • template and catalyzes the formation of new phosphodiester bonds. • This process is repeated as the enzyme moves along the DNA.
RNA Synthesis: addition of new NTPsfollows Watson-Crick rules H N 2 NH 2 NH 2 N O N O N NH N N N HN N N N N O O A•U G•C Template base Incoming base G C C G U A A U
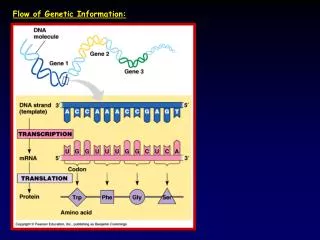
Coding strand 5’…A T G G C C T G G A C T T C A…3’ 3’…T A C C G G A C C T G AA G T…5’ 5’- A U G G C C U G G A C U U C A…3’ Transcription Non-coding (template) strand Translation …Met-Ala-Trp-Thr-Ser… Peptide The sequence of the RNA transcript is complementary to the transcribed DNA strand and is the same as the coding strand
Coding strand 5’…A T G G C C T G G A C T T C A…3’ 3’…T A C C G G A C C T G AA G T…5’ 5’- A U G G C C U G G A C U U C A…3’ Transcription Non-coding (template) strand Translation …Met-Ala-Trp-Thr-Ser… Peptide The sequence of the RNA transcript is complementary to the transcribed DNA strand and is the same as the coding strand DNA RNA
2Pi RNA vs DNA synthesis • Similarities: • • both require a DNA template • • similar mechanism of nucleotide addition • 5’ to 3’ directionality • • Addition of nucleotides follows Watson-Crick base pairing rules • Differences • Only a small portion of one DNA strand is copied • • RNA Pol does not require a primer • • RNA Polymerases do not have exonuclease activity • • RNA Pol is not able to proof read for mismatches • • RNA synthesis is slower than DNA synthesis (50 nt/sec)
The three-dimensional structure of prokaryotic and eukaryotic RNA polymerase Thermus aquaticus Sacchararomyces Cerevisae
E. coli RNA Polymerase Structure • RNA polymerases from bacteria are very large (> 400 kd) composed of four types of subunits: • Subunit Number Mass, kd Function • a 2 37 Binds regulatory sequences • b 1 151 Forms phosphodiester bonds • b' 1 155 Binds DNA template • s 1 70 Recognizes promoter and initiates synthesis • a2bb's = holoenzyme • a2bb‘ = core enzyme
Overview of RNA synthesis in E.coli • 1. Initiation: • • search DNA to find initiation (promoter) sites • • unwind a short region of DNA to expose the template strand • • select correct nucleotides and form the first phosphodiester bond • 2. Elongation • • add NTPs to the growing chain following Watson-Crick • base pairing rules • • locally unwind and re-wind DNA duplex to form transcription bubble • totally processive • 3. Termination • • detect termination signals and pull RNA away from DNA duplex
Initiation signal is present in the 5’ upstream (promoter) sequence of each gene
Consensus Prokaryotic Promoter sequence -10 -35 +1 5' TTGACA TATAAT 3' Coding sequence Promoter sequence Start site • Over 2000 promoter sites in E. coli • strong promoters: 1 transcript/sec • weak promoters: 1 transcript/10 min
subunit recognizes promoter sequences • Roles of subunit: • Decrease the affinity of RNA Pol towards general DNA regions • Enable the recognition of promoter sequence and the initiation of • transcription
E. coli contains multiple factors to recognize several promoter types 70 32 54 Stryer Fig. 28.6
RNA Synthesis 17 base pairs
Initiation of RNA Polymerization O O O G or A -O P O P O P O O- O- O- O 3' OH OH
RNA Synthesis - Continued Transcription bubble
Elongation phase: Transcription bubble Separation of RNA-DNA duplex 50 nt/sec • unwound region 17 bp • RNA-DNA hybrid duplex 8 bp • Mistakes 1 per 104- 1 per 104 nucleotides
RNA Synthesis in Bacteria Start of Gene Several RNA Pol transcribing the same gene
G•C A•U C•G C•G G•C C•G C•G G•C Termination of RNA Polymerization C Two mechanisms possible 1) Rho () protein independent termination stable hairpin formation 2) Rho () protein-mediated termination U G U G A•U•U•U-3' 5'-U•C•C•C•A•C
Antibiotic inhibitors of RNA polymerization Rifampicin inhibits initiation by blocking a channel within RNA polymerase: Actinomycin intercalates in ds DNA, inhibiting strand separation: phenoxazone
RNA synthesis in Eukaryotes See Stryer 5th edition p. 792-798
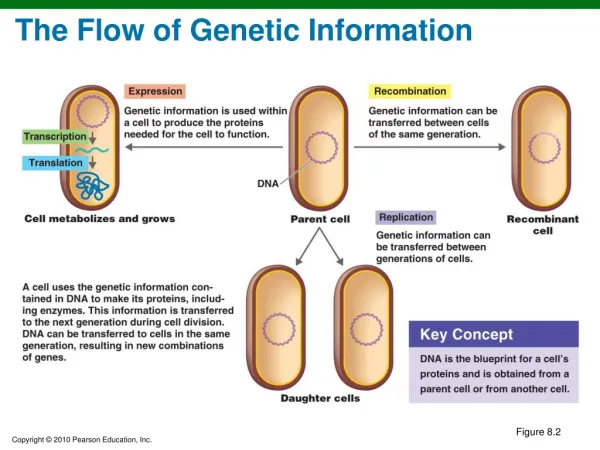
Eukaryotic vs prokaryotic transcription Prokaryotes: • no membrane-bound nucleus • transcription and translation are coupled Eukaryotes: • DNA is located in membrane-bound nucleus • Transcription and translation are separated in space and time
a-amanitin RNA Pol II Inhibitor Amanita phalloides (the death cap)
Actinomycin D •Antitumor antibiotic from Streptomyces genus • Aromatic ring intercalates between GC base pairs, while the peptides bind to the minor groove • Binds to GpC sequences in double-stranded DNA, stabilizing the duplex and inhibiting transcription • Inhibitor of eukaryotic RNA Pol I
Eukaryotic RNA Polymerase II • RNA Pol II is responsible for transcription to pre-mRNA • - 8-12 subunits • - Two large subunits are responsible for RNA synthesis • - Regulated by phosphorylation of carboxyl-terminal domain (CTD) of the largest subunit • - Unphosphorylated form is involved in initiation and phosphorylated form in elengation • - some subunits are shared for RNA Pol I-III
Type II Eukaryotic Promoters -25 +1 -90 -70 5' CAAT Box 3' GC Box TATA box (TATAAAA) (GGNCAATCT) (GGGGCG) Coding sequence Promoter sequence Start site Consensus sequence: Examples:
Initiation of transcription in eukaryotes • RNA Pol II can not initiate transcription by itself • Transcription factors (TFII) are required • The key initiation step is the recognition of TATA box by TBP
Hydrogen bond donors and acceptors on each edge of a base pair
Formation of RNA Polymerase II pre-initiation complex IID contains TBP that binds TATA box IIA stabilizes IID binding to promoter IIB binds initiation sequence Pol II binds IIB IIE stimulates transcription IIH has kinase and helicase activity
Upstream elements (CAAT, GC) Proximal elements (TATA box) Enhancers Coding region Regulated expression Basal promoter Transcriptional control in eukaryotes
Post-transcriptional modifications of mRNA in eukaryotes • 5’ end CAP • polyA tail • splicing
RNA splicing in eukaryotes Primary transcript, hnRNA
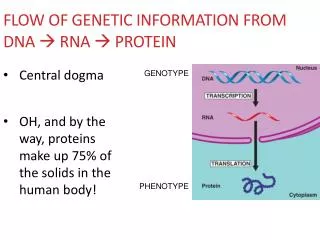
RNA Synthesis: Take Home Message 1) DNA sequences are translated into RNA messages by RNA polymerases. 2) The initiation of RNA synthesis is controlled by specific DNA promoter sequences. 3) The synthesis of RNA is governed by initiation, elongation, and termination steps. 4) Eukaryotic mRNA is extensively processed