Forward Genetics
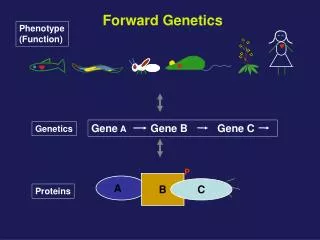
Phenotype (Function). Forward Genetics. Gene A Gene B Gene C . Genetics. P. B. A . C. Proteins. Survey. In classical or forward genetics, we commonly use chemicals or radiation to generate mutations in model organisms such as yeast, worm, fly or mouse.

Forward Genetics
E N D
Presentation Transcript
Phenotype (Function) Forward Genetics Gene A Gene B Gene C Genetics P B A C Proteins
Survey In classical or forward genetics, we commonly use chemicals or radiation to generate mutations in model organisms such as yeast, worm, fly or mouse. In doing so, we typically try to generate mutations in A: somatic cells B: germline cells (gametes, sperm or oocytes) C: not sure
Male XY Sperm X Y 50% 50% XX XY 50% 50% Mutagenesis-sex chromosome female XX 2n meiosis Oocyte Oocyte X X gametes fertilization
xX xY heterozygote mutant Mutagenesis-sex chromosome female Male Mutagen XX XY 2n meiosis Oocyte Oocyte Sperm X x X Y gametes fertilization
Mutagenesis-autosome female Male A A A A 2n meiosis Oocyte Oocyte Sperm y x A A A A gametes 50% 50% fertilization AA AA 50% 50%
X-ray Male A A some Sperm x y A A a AA AA Aa 50% Heterzygous mutant 50% Mutagenesis-autosome female A A 2n meiosis Oocyte Oocyte A A gametes 50% 50% fertilization
Your opinions When you mutagenize the gametes of P0 animals, you usually do not get homozygous mutants in the F1 generation. A: Yes B: No C: not sure. Do you usually get homozygous mutants in the F2? A: Yes B: No C: not sure.
+ + + + Po WT X m + m + F1 WT X m m F2 mutant F1 screens vs. F2 screens + + + + Po WT X m + F1 mutant But, how do we get the same m in two F1 and let them mate?
Balancer chromosomes Chromosomes that suppress crossover. Homozygous of the chromosome is either lethal or with a visible phenotype. Usually contain inversions and translocations A B wt balancer B A A B Resulting abnormal chromosomes B A Neither can be paired with WT
* * X * * F3 homozygotes Traditional F2 screen in fly X-ray TM * X TM TM * X F1 TM
x xX F2 screens in the worm Hermaphrodite XX Po meiosis Sperm Oocyte X gametes X 100% 100% fertilization XX F1 100% Self-progeny
Worm F2 screen xX F1 meiosis Sperm Oocyte x X gametes X X xx 1/4 F2 Homozygous mutant
summary + + + + + + Po WT Po WT X mutagen mutagen + + m + F1 WT X m + F1 WT m + m + F1 WT X m m F2 mutant m m F3 mutant worm Fly, mouse, …
Basic mutagenesis in the worm each F1 carries two mutagenized chromosomes m/+ heterozygotes for each mutation Treat with 50mM EMS for 4 hrs F1 Recovering for a few hours Remove F1 worms after laying ~ 2000 eggs for 20 hrs F2 eggs Several L4 or young adult worms/plate. Multiple large plates Worms with m/m genotype for each mutation are mixed with m/+ and +/+ animals P0 F2 Remove P0 parents after laying ~100 eggs F1 eggs mutant worms (m/m) are individually picked on to a fresh small plate. Total genomes screened = 2X # of F1 animals x # of F1 plates
Question If you plan to mutagenize and screen for a mutation in a tumor suppressor gene that may leads to tumorigenesis, would you do F1 screen or F2 screen? A: F1 B: F2
Physical map YACs cosmids fem-1 skn-1 nhr-48 rme-2 unc-5 eP14 Named genes Genetic and physical map Genetic map nDf41 mDf4 1.8 2.0 2.2 map unit unc-5 skn-1 rme-2 Cloned genes him-3 sup-23 mor-2 unc-77 sup-41 Non-cloned genes evl-7 fem-1 Genetic map and physical map will completely unified when every gene has been mutated. A: yes. B: No. C: not sure.
Pick several male progeny unc unc dpy + X Uncoordinate (Unc) Wild type 50% 50% Pick several wild-type hermaphrodites and place each to one pate + + unc + ; dpy + unc + If the two genes are unlinked ; unc + + dpy unc + + + If the two genes are linked Segregate no dumpy progeny, discard Segregate dumpy progeny, continue mapping with the plates Figure 8.15. Linkage analysis between two mutations. dpy dpy X A few wild type A few Dumpy (Dup)
Dpy-and-Unc 1/16 Unc 3/16 Dpy 3/16 unc + unc + unc unc + + dpy + dpy dpy dpy dpy dpy dpy unc unc dpy + ; ; ; ; ; unc unc + + + + dpy + ; ; 4/16 2/16 2/16 2/16 unc + + + ; 2/16 1/16 1/16 + + + + ; 2/16 1/16 unc + dpy + ; Pick 20 Unc animals If ~ 2/3 of these Unc worm segregate Dpy-and-Unc animals, non-linkage between the two genes is deduced. Situation 1. The two genes are unlinked A single plate Wild type 9/16 Phenotypes of worms In the plate genotypes
unc dpy Wild type 1/2 Unc 1/4 Dpy 1/4 Dpy-and-Unc 0 Phenotypes + dpy unc + + dpy + dpy unc + unc + Non-recombinant Genotypes unc dpy unc dpy unc dpy unc + unc dpy + dpy Rare recombinants Pick 20 Unc animals majority Rare unc dpy unc dpy unc + unc + linkage between the genes is indicated by segregation of no Dpy animals from all or the majority of Unc animals. The frequency of rare recombinants that segregate Dpy animals is correlated with the genetic distance between the two genes. Situation 2. The two genes are linked on one of the six chromosomes + dpy unc + A single plate Self fertilizing
Figure 8.16. Example of genetic three point mapping phenotype genotype dpy unc mapping strain Wild type hermaphrodite egl meiosis Rare gametes from a recombination event Most of the gametes dpy dpy unc egl Sperms or eggs unc Sperms or eggs egl Progeny from combination between common gametes and recombinant gametes Progeny from combination of the common gametes Self-fertilizing unc + dpp + egl dpy Dpy non-Unc unc + dpp + egl + Wild type unc + dpp unc + + Unc non-Dpy unc + dpp unc + dpy Unc and Dpy unc + + + egl + Wild type Egl + egl + + egl + Egl + egl dpy + egl +
dpy unc egl Recombination occurs to the left of egl to the right of the egl dpy dpy egl unc egl unc unc + dpp + + dpy unc + dpp + egl dpy Recognizable recombinants unc + dpp unc egl + unc + dpp unc + + Dpy non-Unc No Unc non-Dpy Yes Dpy non-Unc Yes Unc non-Dpy No Progeny with Egl phenotype unc dpy egl # of recombinations occurred to the left of egl # of recombinations occurred to the right of egl a b = Map position a b
Figure 8.17. An example of genetic mapping using SNPs egl unc X unc egl A C. elegans strain from Hawaii. SNPs between this strain and the Bristol strain have been determined. Genetic mutant derived from the strain from Bristol, England. The egl mutation is being mapped. egl unc The hybrid strain. Stars indicate SNPs in the region X X X * * * * * * 1 2 3 4 5 6 Select Unc but non-Egl recombinants B A C egl unc egl unc egl unc * * * * * * * * * * * * * * * * * * unc unc 2 3 4 5 6 unc 4 5 6 6 Determine SNP #4 for all recombinant worms by sequencing or digestion. Worms A and B have #4 SNP from the Hawaii strain Determine SNP #5 and #6 for those that have lost SNP#4 (worm C only) Worm C has SNP #6 but not #5: the egl gene maps to the right of SNP#5
Mapping using marker mutations SNP mapping Microinjection of cosmid/YAC clones RNAi of candidate genes Injection of subclones sequencing mutant DNA Common steps involved in cloning C. elegans genesdefined by mutations. Genetic mutation What would be the flow chart for cloning in yeast? Fly? Human?
Which is the strongest evidence for claiming the cloning of the gene defined by the mutation? A. The transgene put back into the animal can rescue the mutant phenotype. B. You find a missense mutation in this gene by sequencing. C. Reducing the gene activity by RNAi mimics the mutant phenotype. D. The gene is expressed in the tissue with the mutant phenotype.
Figure 8.19. Microinjection transformation in C. elegans. DNA solution is injected to the distal arms of the gonad Injection Select F1 transgenic animals, most are unstable F2 transgenic animals, stable lines Three types of markers unc-119(-) mutant Wild type Wild type a strongly expressed GFP gene as a marker unc-119(+) gene as a marker A dominant rol-6 mutant gene as a marker Wild type Roller Green worm
Figure 8.21. The prevailing model for the mechanism of RNAi dsRNA Introduced into cells Dicer Bind to Dicer-RDE-1enzyme complex dsRNA is cut to ~22 nt siRNA Incorporated into RISC nuclease complex Multiple-protein components of RISC Unwinding siRNAs, activation of RISC Target mRNA AAAAAAAA 5’ Cleavage of target mRNA
Figure 8.22. RNAi methods in C. elegans. A. Injecting ds RNA into intestine or gonad dsRNA intestine Transfer to plates Observe phenotype in progeny B. Soak worms with dsRNA solution Soak for 24 hours Transfer to plates Observe phenotype in progeny C. Feed the worms a bacterial strain that expresses dsRNA Feed worms the bacterial strain Grow the bacterial strain containing the vector expressing dsRNA Observe phenotype in progeny