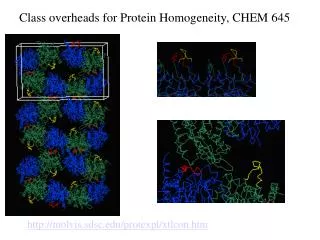
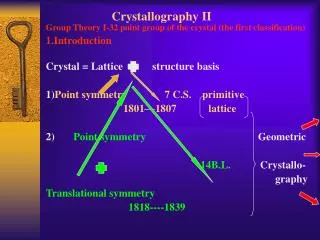
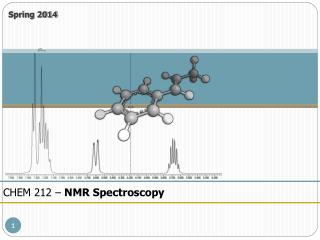
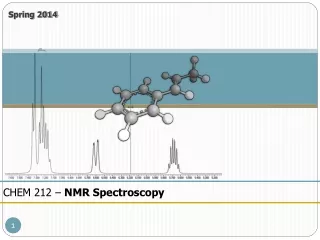
NMR vs. Crystallography for CHEM 645
NMR vs. Crystallography for CHEM 645. Brian Bahnson Department of Chemistry & Biochemistry University of Delaware. Distance Restraints. Through space - NOE. NOE 1/r 6 . f ( t c ). Tortional Restraints through bond J-coupling. NMR Refinement. ideal geometry. NMR term.

NMR vs. Crystallography for CHEM 645
E N D
Presentation Transcript
NMR vs. Crystallographyfor CHEM 645 Brian Bahnson Department of Chemistry & Biochemistry University of Delaware
Distance Restraints Through space - NOE NOE 1/r6.f (tc)
Tortional Restraints through bond J-coupling
NMR Refinement ideal geometry NMR term = wNMR distance restraint + tortional restraint + wideal Etotal violations violations X-ray Refinement ideal geometry X-ray term = wFwhkl (|Fo| - |Fc|)2hkl + wideal Etotal hkl calculated observed
13C, 15N labeling, homogeneity Bigger magnet is better – 600, 750 or 900 MHz 2-D and 3-D homonuclear and heteronuclear pulse sequences Ikura et al., (1989) Biochemistry 29, 4659-4667., then do side chains NOE: Wuthrich, (1989) Science 243, 45-50. also: J-coupling ~ tortion Pattern recognition, build 100 models, select 20 best Minimize restraint violations, keep “good” geometry = WN(distance restraint violation + WI (Etotal)
NMR Structures of closed form calmodulin
X-ray Crystal Structures of calmodulin
Bundle of 20 NMR models of calmodulin • Cases of Bundle Spread • Missing restraints • dynamics
Crystallography vs. NMR – advantage/disadvantages • Experimental difficulties • need for homogeneity in common • need good crystals for crystallography • need 13C and 15N label for NMR • size limits of NMR technique • solubility an issue for each technique • Reported structure(s) look different – i.e. bundle • crystal vs. solution structure • Complementary information • high resolution vs. dynamics • positional amplitude, • certainty time domains
Molecular Replacement – homology modeling Molecular Replacement (MR) – another method to estimate phases – use a structurally homologous protein >25% sequence identity is sometimes possible >50% sequence identity is a safe bet • Make search model • - find structural model of sequence homolog • - from sequence alignment and homolog structure, create model • - mutate or trim down to what the two proteins have in common • - energy minimize to eliminate bad geometry (intro to refinement) Suppose you wanted to make a model of BSIDH
Homology Modeling Links Swiss Model http://swissmodel.expasy.org//SWISS-MODEL.html NCBI PubMed http://www.ncbi.nlm.nih.gov/sites/entrez/ Homology modeling tutorial http://molvis.sdsc.edu/protexpl/homolmod.htm Principles of Protein Structure, Comparative Protein Modeling and Visualization (http://swissmodel.expasy.org//course/course-index.htm)DeepView - download a free version of this viewer. Its also for linux computers. (http://au.expasy.org/spdbv/) A tutorial for Deep View was made by Gale Rhodes, the author of CMCC. (http://www.usm.maine.edu/~rhodes/SPVTut/index.html)