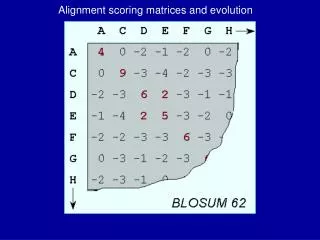
Scoring Matrices
Scoring Matrices. Diff. Scoring Rules Lead to Diff. Alignments. Example Score = 5 x (# matches) + (-4) x (# mismatches) + + (-7) x (total length of all gaps) Example Score = 5 x (# matches) + (-4) x (# mismatches) + + (-5) x (# gap openings) + (-2) x (total length of all gaps).

Scoring Matrices
E N D
Presentation Transcript
Diff. Scoring Rules Lead to Diff. Alignments • Example Score = 5 x (# matches) + (-4) x (# mismatches) + + (-7) x (total length of all gaps) • Example Score = 5 x (# matches) + (-4) x (# mismatches) + + (-5) x (# gap openings) + (-2) x (total length of all gaps)
Scoring Rules/Matrices • Why are they important? • The choice of a scoring rule can strongly influence the outcome of sequence analysis • What do they mean? • Scoring matrices implicitly represent a particular theory of evolution • Elements of the matrices specify the similarity of one residue to another
The alignment score of aligning two sequences is the log likelihood ratio of the alignment under two models • Common ancestry • By chance
Likelihood Ratio for Aligning a Single Pair of Residues • Above: the probability that two residues are aligned by evolutionary descent • Below: the probability that they are aligned by chance • Pi, Pj are frequencies of residue i and j in all sequences (abundance)
Two classes of widely used protein scoring matrices PAM = % Accepted Mutations:1500 changes in 71 groups w/ > 85% similarityBLOSUM = Blocks Substitution Matrix:2000 “blocks” from 500 families
PAM and BLOSUM matrices are all log likelihood matrices • More specifically: • An alignment that scores 6 means that the alignment by common ancestry is 2^(6/2)=8 times as likely as expected by chance.
Constructing BLOSUM Matrices Blocks Substitution Matrices
BLOSUM Matrices of Specific Similarities • Sequences with above a threshold similarity are clustered. • If clustering threshold is 62%, final matrix is BLOSUM62
A toy example of constructing a BLOSUM matrix from 4 training sequences
BLOSUM matrices reference • S. Henikoff and J. Henikoff (1992). “Amino acid substitution matrices from protein blocks”. PNAS 89: 10915-10919 • Training Data: ~2000 conserved blocks from BLOCKS database. Ungapped, aligned protein segments. Each block represents a conserved region of a protein family
Break • Homework
PAM Matrices (Point Accepted Mutations) Mutations accepted by natural selection
Total Mutation Rate is the total mutation rate of all amino acids
Normalize Total Mutation Rate to 1% This defines an evolutionary period: the period during which the 1% of all sequences are mutated (accepted of course)
Mutation Probability Matrix Normalized Such that the Total Mutation Rate is 1%
-- PAM1 mutation prob. matr. -- PAM2 Mutation Probability Matrix? -- Mutations that happen in twice the evolution period of that for a PAM1
In two PAM1 periods: • {AR} = {AA and AR} or {AN and NR} or {AD and DR} or … or {AV and VR}
PAM60—60%, PAM80—50%, • PAM120—40% • PAM-250 matrix provides a better scoring alignment than lower-numbered PAM matrices for proteins of 14-27% similarity
PAM Matrices: Reference • Atlas of Protein Sequence and Structure, Suppl 3, 1978, M.O. Dayhoff. ed. National Biomedical Research Foundation, 1
PAM Based on extrapolation of a small evol. Period Track evolutionary origins Homologous seq.s during evolution BLOSUM Based on a range of evol. Periods Conserved blocks Find conserved domains Comparing Scoring Matrix