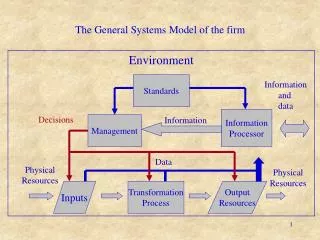
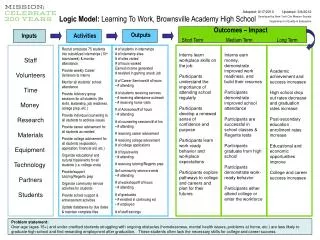
What (inputs)
my Grid an e-biologist’s workbench www.mygrid.org.uk candidate genes Gene annotation pipeline - what is known about my gene? Provenance of e-Science Experiments experience from bioinformatics

What (inputs)
E N D
Presentation Transcript
myGrid an e-biologist’s workbench www.mygrid.org.uk candidate genes Gene annotation pipeline - what is known about my gene? Provenance of e-Science Experiments experience from bioinformatics The myGrid project aims to provide middleware layers that make the Information Grid appropriate for the needs of bioinformatics. Provenance management is one of the services provided to support scientific method and best practice found at the bench but often neglected at the workstation. Provenance metadata records the materials, methods and results of in silico experiments as well as scientists' notes about experiments. This information is essential if experiments are to be validated and verified by others. It is also important in assessing the quality and timeliness of results. The large amount of provenance data required to verify others' experiments means that as much as possible must be automatically and systematically collected. Sets of provenance data provide a store of know-how that can be mined to provide users with views of their virtual organisation, including suggestions based on past experience. Bioinformatics analyses typically involve visiting many data resources and analytical tools. These in silico experiments can be created as pipelines or “workflows” in the Taverna editor. Claire Jennings Gene ID 504_at 2003/08/15 20:25 who What (inputs) Provenance metadata includes standard annotations, e.g. who performed the in silico experiment and when. Scientists can also provide additional context information such as hypotheses, conclusions and related literature when what The provenance record of a workflow run is a convenient way of automatically and systematically recording the derivation path by which results are generated from input data. The workflow, services and input data in a workflow provenance record are all items which have their own provenance how why Services What (outputs) The use of semantically rich provenance metadata expressed using ontologies enables the creation of a “web of scientific holdings” where items are linked by related concepts Acknowledgements This work is supported by the UK e-Science programme EPSRC GR/R67743, & DARPA DAML subcontract PY-1149, Stanford University. The Graves Disease case study is from Simon Pearce and Claire Jennings, Institute of Human Genetics, School of Clinical Medical Sciences, University of Newcastle. The myGrid team: Matthew Addis, Nedim Alpdemir, Rich Cawley, Vijay Dialani, Alvaro Fernandes, Justin Ferris, Rob Gaizauskas, Kevin Glover, Carole Goble (director), Chris Greenhalgh, Mark Greenwood, Ananth Krishna, Phil Lord, Xiaojian Liu, Darren Marvin, Karon Mee, Simon Miles, Luc Moreau, Tom Oinn, Juri Papay, Norman Paton, Steve Pettifer, Milena Radenkovic, Peter Rice, Angus Roberts, Alan Robinson, Martin Senger, Nick Sharman, Robert Stevens, Paul Watson, Anil Wipat & Chris Wroe.